Documente Academic
Documente Profesional
Documente Cultură
Healthcare Insulin Problem
Încărcat de
Taufiq JaiDrepturi de autor
Formate disponibile
Partajați acest document
Partajați sau inserați document
Vi se pare util acest document?
Este necorespunzător acest conținut?
Raportați acest documentDrepturi de autor:
Formate disponibile
Healthcare Insulin Problem
Încărcat de
Taufiq JaiDrepturi de autor:
Formate disponibile
{
problem # 6
Manufacturer of Insulin
Problem submitted by: Professor David Meredith, P. E., Pennsylvania State UniversityFayette, Uniontown, PA
Problem Statement
Bioengineers are responsible for developing processes such as the recombinant DNA process described here. They also develop articial joints and human organs and medical tools to support surgeons. Chemical engineers are involved with sizing the process equipment and developing the ow path through the process. Electrical and mechanical engineers build the support systems such as the instrumentation and distilled water supply. To meet this years increase in global demand, your team is to design a manufacturing facility capable of producing 1,800 kg/yr of human insulin. You will be using a recombinant DNA process by growing E. coli bacteria that contain Trp-LE-Met-proinsulin. You will need to provide the proper growth media, air, heat and agitation. After a batch has matured, you must homogenize the material then separate the product using centrifuges, dialters and chromatography in a series of steps with specic conditions as you extract and purify the product. Finally, the insulin crystals must be freeze-dried into crystals for nal distribution.
Background
Human insulin is a polypeptide containing 51 amino acids arranged in two chains. The A chain contains 21 amino acids and the B chain includes 30 amino acids. The two chains are linked by two disulde bonds. Human insulin has a molecular weight of 5,734 and an isoelectric point of 5.4. In the proinsulin process (see Figure 6-1), the plasmid circle of DNA (1) that exists in bacteria are genetically modied to include part of the human DNA (2). This recombinant plasmid (3) is inserted into Eschericia coli (E. coli) (4). A large colony of the inoculated E. coli bacteria cells are fermented to overproduce Trp-LEMet-proinsulin in the form of inclusion bodies (5), which are recovered and soluablized. Proinsulin is released by cleaving the methionine linker using CNBr (6). The proinsulin chain is subjected to a folding process to allow intermolecular disulde bonds to form (7), and the C peptide is then cleaved with enzymes (8) to yield human insulin (9). (Note the process described here has been simplied and several key steps have been omitted. Dont try this at home.)
Figure 6-1 Production Process
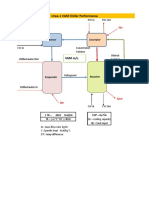
The rst step of the process is fermentation ( See Figure 6-2 ). The nutrients are premixed and added to deionized water in a large tank with an agitator. When conditions are acceptable, a seed colony of inoculated bacteria are added to the tank. The tank temperature is maintained at 37C by a water jacket that can be either heated (early in the cycle when heat loss from the reactor is greater than the heat produced by the bacteria) or cooled (later in the cycle when the bacteria produce more heat than the reactor loses).
weight per liter of broth. The basic unbalanced chemical equation is:
C6H12O6 + H2O + NH3 + (seed bacteria) => CH1.8O0.5N0.2
(6-1)
Atomic masses for selected elements are given below: Hydrogen=1.008 Nitrogen=14.007 Sulfur=32.064 Bromine=79.91 Carbon=12.01 Oxygen=15.999 Zinc=65.37
Assumptions and Givens
The manufacturing facility operates around the clock for 330 days per year. A new batch is initiated every 48 hours resulting in 160 batches per year. This implies a required cycle production of about 11.3 kg of insulin crystals per batch. See Figure 6-2. A cubic meter is 1,000 liters and a cubic centimeter (cm3) is equal to one ml. A micron is 10-6 meter. Each batch requires glucose (C6H12O6) to feed
51. The mass of glucose (kg) required to grow one batch of the e. coli bacteria to the nal concentration is closest to: a. 1,350 b. 7,500 c. 4,500 d. 18,500 e. 225,000
the bacteria (CH1.8O0.5N0.2) in 30 m3 of broth to grow to a nal concentration of 37 g of dry cell
Additional Assumptions and Givens
In addition to temperature and food (glucose), the bacteria also need oxygen to grow. The oxygen is
Nutrient Premix Tank Agitator
Compressor
Air Filter
Temperature Control Jacket
To Cell Recovery
Figure 6-2 Fermentation Process
provided by air, which is 21% oxygen by volume. The air is compressed by a 50 kW compressor to 7 atmospheres of pressure and is ltered through a cartridge air lter before being sparged (released as tiny bubbles) into the bottom of the reactor. As the air bubbles rise through the broth, some of the oxygen dissolves into the broth solution to be taken up by the bacteria. The bacteria require 0.35 g of O2 per g of living cells each hour. The minimum dissolved oxygen concentration to support growth is 0.2 mg / liter of broth. The maximum dissolved oxygen concentration in the medium is 6.7 mg/liter at 37C. The amount of oxygen required to support the colony growth is given by:
The agitator is 2.5 m in diameter and stirs the tank at 60 revolutions per minute. The motor input power, Po (kW), required by this agitator is given by:
Po = 0.7 N3 Da2 V
where,
(6-3)
N = speed of the agitator (revolution per second) Da = diameter of the agitator (m) V = the volume of the broth in the reactor (m3)
Because aerated mixtures are less dense than nonaerated mixtures, the motor power is reduced. The input, Pg (kW), for the agitator for the bioreactor is empirically given by:
3 � Po2 NDa 0.56 � Q � 0.45
k (C CDO ) X= LA DO
where,
qO 2
(6-2)
� Pg � k � � where,
(6-4)
L = liter X = nal concentration of cells (dry cell weight) per liter of broth (g/L ), kLA = oxygen volume transfer rate (m3 O2 / m3broth-h) CDO* = maximum dissolved oxygen (DO) concentration in the broth (gO2/L) CDO = minimum DO concentration to support growth (gO2/L) qO2 = respiration rate of cells (gO2/gcell h)
52. The ow rate of air (m3/s) to maintain the broth at the nal cell concentration is closest to: a. 2.3 d. 81 b. 17 e. 292 c. 70
k = proportionality constant (use 0.5) Q = volumetric ow rate of gas per volume of tank (s-1)
53. When the volumetric ow rate of gas is equal to 15 (s-1), the electrical input (kW) to operate the agitator is closest to: a. 71 d. 813 b. 97 e. 2,268 c. 140
Additional Assumptions and Givens
At the end of the 18-hour fermentation time, the broth is cooled from 37C to 10C to quench the biological action. This cooling process occurs in about 30 minutes. Chilled water at 5C is injected into the reactor jacket at a ow rate of 50 L/s, picks up heat from the broth and exits the reactor jacket to return to the cooling system. The density and specic heat of the broth are 1,020 kg/m3 and 3.8 kJ/kg-C respectively. The density and specic heat of the water are 1,000 kg/m3 and 4.19 kJ/kg-C respectively.
Additional Assumptions and Givens
To maintain constant growing conditions throughout the broth, the aerated mixture is stirred continuously by a large agitator that looks like a kitchen mixer on steroids.
The energy balance equation is given by:
From Fermentation
(V
cp T )broth = (Q t cp T )water
(6-5)
where,
Holding Tank
Homogenizer To Solubilization Centrifuge Centrifuge
cp = specic heat of the uid (kJ/kg-C) = density of the uid (kg/m3) V = volume of the broth (m3) Q = volumetric ow rate of chilled water (m3/h) t = time duration of chilled water ow (h) T = temperature difference for the uid before and
after the heat transfer occurs (C) 54. The average water temperature in (C) leaving the reactor jacket during this cooling period is closest to: a. 6 d. 20 b. 8 e. 35 c. 13
Figure 6-3b. Cell Recovery Process
The structure of EDTA is shown above in Figure (6-3a) 55. The molecular weight of EDTA is closest to: a. 265 d. 292 b. 276 e. 340 c. 286
Additional Assumptions and Givens
A high-pressure homogenizer is used to break the cells and release the inclusion bodies. Passing the 15,000 liters of buffered sludge through the homogenizer three consecutive times through a circular nozzle at high velocity and under a pressure change of 80 MPa each time, ruptures the cell walls to expose the inclusion bodies. The homogenizer capacity is 10,102 L/h and the electrical input to the motor is 225 kW. It takes about 1.5 hrs for each pass or 4.5 hrs total for the process. The density of the sludge at this point is 985 kg/m3.
Additional Background
Once the broth has been cooled, it is transferred to a holding tank to allow the fermentation tank to be cleaned and prepared for the next cycle. To reduce the volume of material to be processed, the 30,000 liters of broth are passed through a centrifuge to reduce the volume by a factor of four. This process also removes most of the extra-cellular impurities. Next the thickened cell sludge is diluted with an equal volume of buffer solution consisting of 94.4% distilled water, 0.7% EDTA (ethylenediaminetetraacetic acid), a sequestering agent to prevent further reactions and 2.9% TRIS-Base (trishydroxymethylaminomethane), a buffering agent. This process facilitates the separation of the cell debris particles from the inclusion bodies that contain the proinsulin.
Additional Assumptions and Givens
The pressure change, P (kPa), is given by: (6-4) 2 P = V / 2,000 where, = density of the uid (kg/m3) V = throat velocity through the nozzle (m/s) The ow rate of uid, Q (m3/s) is given by:
Q = VA
where, Figure 6-3a EDTA Structure
(6-5)
V = throat velocity (m/s) A = cross-sectional area of the nozzle throat (m2)
56. The nozzle diameter (mm) is closest to: a. 3 d. 17 b. 5 e. 22 c. 9
= kinematic viscosity (m2/s) of the uid = shape factor (use 1.0 for spherical)
The process time, t (s) is given by:
Additional Assumptions and Givens
The 15,000 liters of homogenized sludge is returned to the centrifuge to separate the inclusion bodies from the cell debris. The inclusion bodies have a diameter of 1.0 microns and a density of 1.3 g/cm3. Assume water ( = 1.0 g/cm3) is the liquid media with a kinematic viscosity of 1.004x10-6 m2/s. The centrifuge has an acceleration factor of 1,500g (one g is equal to an acceleration of 9.81 m/s2), has 147,718 m2 of effective clarifying surface area and a motor input of 25.6 kW with a yield of 98%. The centrifuge reduces the volume from 15,000 liters to 1,400 liters of slurry at the end of the recovery section. The throughput or ow rate of entering uids to be separated, � (kg/s) is given by:
2 2
t = where,
(6-7)
Q = entering ow volume to be separated (m3) = density of the liquid media entering the lter (kg/m3) � = throughput (kg/s)
57. The length of time (h) required to process each batch is closest to: a. 1.8 d. 8.3 b. 2.3 e. 46 c. 5.3
Additional Background
The 1,400 L of inclusion body suspension is transferred to a glass-lined agitator tank and mixed with small quantities of urea and of 2-mercaptoethanol. Urea is a chaotropic agent that dissolves the denatured protein in the inclusion bodies and 2-mercaptoethanol is a reductant that reduces disulde bonds. It takes a reaction time of 8 hours to solubilize 95% of the proinsulin. The next step is to remove the excess urea and 2-mercaptoethanol with a dialter and replace it with distilled water. This solution is then ltered through a dead end lter to remove ne particles that might overload ltering processes downstream. See Figure (6-4).
d (s l )(r )(V D) Fs = 18 where,
(6-6)
d = particle diameter (m) s = density of the particle (g/cm3) 1 = density of the uid media (g/cm3) (r2) = acceleration factor (m/s2) V / D = effective clarifying surface area (m2) Fs = correction factor for fraction of solid present (use
0.05 for 20% solids) From Cell Recovery Distilled Water Agitated Tank Holding Tank
Additional Assumptions and Givens
The design ux capacity of the membrane used is 4.2 liters per hour for each square meter of membrane.
Filter
To CNBr Cleavage
Diafilter
Reactant
Figure 6-4 Solubilization Process
The dialtration process takes 6 hours to complete. The sizing equation for a dialter is given by:
and excess reactants are removed by applying a vacuum and raising the temperature to 35C (the boiling temperature of CNBr at that pressure). Assume the absolute pressure is 0.28 atmospheres at 35C and you have to boil off 20 kg of CNBr vapor. One atmosphere (atm) is 101 kPa and assume the Ideal Gas law: where,
Q where,
= J A t
(6-7)
Q = volume of media to be processed (liters) J = Design ux capacity (l/m2-h) A = membrane area (m2) t = process time (h)
58. The membrane area (m2) required for this process is closest to: a. 0.02 d. 55 b. 18 e. 590 c. 41
PV = nRT
(6-8)
R = universal gas constant and is 8.314 kPa-m3/[kmol-K] P = pressure in kPa V = volume in m3 n = number of kilomoles of gas T = temperature in degrees Kelvin.
59. The minimum volume (m3) of the CNBr vapor produced at evaporator conditions is closest to: a. 1.940 d. 54.3 b. 9.570 e. 1,800 c. 17.0
Additional Background
The chimeric protein is cleaved in a well-mixed reactor with CNBr (cyanogen bromide) in a 70% formic acid solution into the signal sequence Trp-LE-Met, which contains 121 amino acids and the denatured proinsulin (82 amino acids). The reaction takes 12 hours at 20C. The nal mass of proinsulin is 31% of the mass of the initial Trp-LE-Met-proinsulin. See Figure (6-5). Additional Assumptions and Givens A small amount of toxic cyanide gas is formed as a byproduct of the cleavage reaction. This toxic gas Vacuum
Additional Background
A sultolysis step is used to unfold the proinsulin, break any disulde bonds and add SO3 moieties (molecule segments) to all sulfur residues on the cysteines. This reaction occurs in a well-stirred reactor under alkaline conditions (pH 9 to 11) over a 12-hour period. The reagents are ltered through a dialter and replaced with a 20% by weight solution of HCl guanidine.
HCI + Guanidine Evaporator
Agitated Tank Agitated Tank CNBr Cleavage
Holding Tank
Diafilter Sulfitolysis
Reactant
Figure 6-5 Cleavage and Sultolysis
From Sulfitolysis
Acid Regeneration Filtered solution To Folding Proinsulin solution
Figure 6-6 Ion Exchange Chromatography The human proinsulin(S-SO3-)6 is next chromatographically puried in an ion-exchange column. The bed consists of small beads coated with a cation (group of atoms carrying a positive electric charge) exchange resin. An ionic bond is temporarily formed with the properly formed proinsulin(S-SO3-)6 molecules as the solution passes slowly up through the resin. The molecules are released when the column is regenerated (back-ushed) with acid. See Figure 6-6. hours and reaches a yield of 85%. The reactions are removed and the protein solution is concentrated with a dialtration unit followed by purication in a hydrophobic interaction chromatography (HIC) column to remove unfolded or incorrectly folded molecules. The nal reaction step is to enzymatically remove the C peptide from the human proinsulin using trypsin and carboxypeptidase B. The reaction occurs in a well-mixed reactor at 10C for 12 hours. The reagents are removed by another dialtration step followed by purication in another ion-exchange chromatography (IEC) column. Several additional purication steps are included in the process. A reversed-phase, high-pressure liquidchromotography (PR-HPLC) step removes structurally similar insulin-like components. A dialtration process removes the reactants and concentrates the solution by a factor of two. The nal purication step is a gel ltration chromatographic column followed by another dialtration process to concentrate the solution by a factor of ten. The 500-liter solution at this point contains 12.8 kg of insulin. An ultralter captures particles larger than 100 nanometers (>10-7 m). See Figure 6-7. The nal step is to crystallize the nal insulin. The insulin solution is mixed with ammonium acetate and zinc chloride in an agitated reaction tank. The crystallization into insulin6-Zn2 is carried out at 5C for twelve hours. IEC RP-HPLC Gel Filter Agitated Tank Holding Tank Distilled Water Final Purification Diafilter Reactant Enzymatic Conversion Diafilter Figure 6-7 Refolding and Enzymatic Conversion Reactant
Additional Assumptions and Givens
The ion exchange column measures 140 cm in diameter with a bed height of 25 cm. The 8,000 liters of solution at this point contains 28 kg of proinsulin(S-SO3-)6. The maximum holding capacity of the resin is 18 mg of proinsulin(S-SO3-)6 for each ml of nominal bed volume. The regeneration cycle begins when the bed reaches 70% of maximum holding capacity.
Additional Background
Folding of the proinsulin(S-SO3-)6 and disulde bond formation takes place in a a well-mixed reactor using mercaptoethanol to facilitate the disulde interchange reaction. The reaction is carried out at 8C for 12 Distilled Water Agitated Tank Refolding Holding Tank
The 11.3 kg of insulin crystals are separated in a basket centrifuge and freeze-dried at 20C under a vacuum (0.0015 atmospheres).
60. The number of diabetics supported by the insulin produced at this facility is closest to: a. 2,100 d. 1.8x106 b. 4,200 e. 4x106 c. 11,500
Additional Assumptions and Givens
An average diabetic individuals daily insulin usage is 60 units. (This quantity varies with body weight, activity level and diet.) A unit is 1/22 mg of crystallized insulin.
Competition
Part I and Part II Solutions 2007
Problem #6 Manufacture of Insulin
51.
Answer a The molecular weight of glucose is: (C 6 H 12 O 6 ) = (6x12) + (12x1) + (6x16) = 180 The molecular weight of the bacteria is: (CH 1.8 O 0.5 N 0.2 ) = (1x12) + (1.8x1) + (0.5x16) + (0.2x14) = 24.6 Six moles of bacteria require one mole of glucose, or 6x24.6 = 147.6g of bacteria require 180g of glucose. Each batch requires enough glucose to feed the bacteria in 30 m 3 of broth to a concentration of 37g of dry cell weight per liter of broth. (37g/lt) (30x10 3 lt) = 1,110 kg of bacteria. This requires: 1,110kg of bacteria x 180g of glucose = 1,353 kg of glucose 147.6g of bacteria
52.
Answer d Using equation (6-2), and solving in terms of K LA yields: K LA = (X)(q02 ) C*DO C DO
where, K LA = Oxygen Volume Transfer (in m 3 02 /m 3 broth - h) X = final concentration of cells = 37g cells /L q 02 = Respiration rate of cells = 0.35g 02 /g cell h
TEAMS 2007 Part I and Part II Solutions
37
C DO = Minimum DO concentration to support growth = 0.2mg 02 / L C * DO = Maximum DO concentration in the broth = 6.7mg 02 / L K LA = (37g cell /L)(0.35g02 /g cell h)(1,000mg02 /g 02 ) (6.7mg02 /L 0.2mg 02 /L)
= 1,992 m 302 /m 3broth h Each batch is 30m 3 and the O2 represents 21% of the air: Also there are 3,600 s/h. Therefore the Volumetric rate of air required is:
(30m3 /batch)(1,992m 302 /m 3broth h) = 79.04m 3air /s 3 3 (3,600s/h)(0.21m 02 /m air )
53.
Answer a Using equation (6-3) with: N = Speed of the Agitator = 60 rev/min = 1 rev/s D a = Diameter of the Agitator = 2.5m, and V = Broth volume in the reactor = 30m 3 , yields: 0 = (0.7)(1rev/s) 3 (2.5m) 2 (30m 3 ) = 131.25 kW Then using equation (6-4) with: k = 0.5 0 = 131.25 kW N = 1 rev/s D a = 2.5m, and Q = 15/s, yields:
TEAMS 2007 Part I and Part II Solutions
38
(131.25 kW) 2 (1rev/s)(2.5m) 3 g = (0.5) (15/s) 0.56
0.45
= 70.2 kW 54. Answer c
broth
= Density of broth = 1,020 kg/m 3
C pbroth = specific heat of broth = 3.8 kJ/kg- 0 C
T
broth
= 27 0 C
water
= 1,000 kg/m 3
C pwater = 4.19 kJ/kg 0 C Q = volumetric flow rate of chilled water (in m 3 /h) = 50 L/s = (50L/s)(3,600s/h) (1,000L/m 3 )
= 180 m 3 /h t = 30min = 0.5h 60min/h
Solving equation (6-5) in terms of T water yields:
Twater = ( broth )(30m3 )(Cpbroth )(Tbroth ) (Q)( water )(Cpwater )(t)
(1,020kg/m 3 )(30m 3 )(3.8kJ/kg 0 C)(27 0 C) (180m 3 /h)(1,000kg/m 3 )(4.19kJ/kg 0 c)(0.5h)
= 8.32 0 C Water Temperature = 5 0 C + T = 13.32 0 C
TEAMS 2007 Part I and Part II Solutions
39
55.
Answer d The molecular formula for EDTA is: C 10 H 16 O 8 N 2 and the molecular weight of the molecule is: (12g x10) + (1g x16) + (8x16g) + (2x14g) = 292 g/mol
56.
Answer a Using equation (6-6) and solving it in terms of V yields: V=
(P)(2,000)
where,
= Pressure Change = 80 MPa = 80x10 3 kPa
= density of fluid = 985 kg/m 3 therefore, V= (80x10 3 kPa)(2,000) = 403.03 m/s 985kg/m 3
Using equation (6-7) and solving it in terms of A yields: A = Cross-Sectional Area of Nozzle Throat
15,000L/h Volume 1,000L/m 3 Q= = Time (90min)(60s/min)
= 2.7x10 3 m 3 /s A= Q 2.7x10 3 m 3 /s = = 6.892x10 6 m 2 V 403.03m/s
A = 6.892 mm 2
TEAMS 2007 Part I and Part II Solutions
40
A=
D 2 , or 4 4A = (4)(6.892mm 2 )
D=
D = 2.96 mm 57. Answer b d = particle parameter = 1.0 microns = 1x10 6 m s = density of the particle = 1.3g/cm 3 1 = density of fluid media = 1g/cm 3 n = kinematic viscosity of fluid = 1.004x10 6 m 2 /s rw 2 = acceleration factor = 1,500g = 1,500 x 9.81m/s 2 V/D = effective clarifying surface area = 147,718m 2
= shape factor = 1.0
F s = correction factor for fraction of solid present = 0.05 Using equation (6-8) yields: (1x10 6 m) 2 (1.3g/cm 3 1.0g/cm 3 )(1,500x9.81m/s 2 )(147,718m 2 )(0.05) = (18)(1.004x10 6 m 2 /s)(1.0) = 1.8 kg/s Using equation (6-9) yields: Q = (15,000 lt)(1 m 3 /1,000 lt) = 15m3 = 1.0g/cm 3 = 1.0x10 3 kg/m 3
t = (15m 3 )(1.0x10 3 kg/m 3 )` = 8,333s = 2.3h 1.8kg/s
TEAMS 2007 Part I and Part II Solutions
41
Note: In reality the density of the entering sludge is 13,600 lt at a density of 1.0g/cm 3 plus 1,400 lt at a density of 1.3g/cm 3 for a total of 15,420 kg in 15,000 lt for a density of 1.03g/cm 3 . 58. Answer d Solving equation (6-10) in terms of A yields: A= Q , where (J)(t)
Q = volume of media to be processed = 1,4000 l J = Design flux capacity = 4.2 l/m 2 -h t = process time = 6h A= 1,400 l (4.2 l/m 2 h)(6h)
= 55.55 m 2 59. Answer c In order to use equation (6-11) we need to calculate (n) the number of kilomoles of gas. The molecular weight of the CNBr vapor is: 12g + 14g + 80g = 106g/mol = 106 kg/kmol The removal of 20 kg represents n= 20kg = 0.189 kmol of gas. 106kg/kmol
Solving equation (6-11) in terms of V yields: V= (n)(R)(T) P
TEAMS 2007 Part I and Part II Solutions
42
(0.189kmol)(8.314kPa m3 /kmolo K)(35 + 273 o K) (0.28atm)(101kPa/atm)
= 17.11 m 3 60. Answer d Facility production = (1,800kg/yr)(1yr/365 days)(10 6 mg/kg) = 4,931,506 mg/day Average daily dose in mg 1 mg = (60 units/day) 22 unit = 2.73 mg/daily dose Number of supported diabetics = 4,931,506mg/day 2.73mg/day
= 1,806,412 diabetics = 1.8x10 6 diabetics
TEAMS 2007 Part I and Part II Solutions
43
S-ar putea să vă placă și
- Distillation Version 3Document4 paginiDistillation Version 3Toru Lucis CaelumÎncă nu există evaluări
- Fermentation (Industrial) : Basic ConsiderationsDocument12 paginiFermentation (Industrial) : Basic ConsiderationsRudi TabutiÎncă nu există evaluări
- PDF Solution of Treybal DLDocument2 paginiPDF Solution of Treybal DLaliÎncă nu există evaluări
- Biochemical Engineering Aiba HumphreyDocument4 paginiBiochemical Engineering Aiba HumphreyYork De Moya LlanosÎncă nu există evaluări
- Specification Sheet For Crystallizer Crystallizer Data SheetDocument3 paginiSpecification Sheet For Crystallizer Crystallizer Data SheetAmar RazinÎncă nu există evaluări
- Volumen Molar Parcial, Densidad y ViscosidadDocument8 paginiVolumen Molar Parcial, Densidad y ViscosidadMichelle CastilloÎncă nu există evaluări
- Tabla 5B APIDocument24 paginiTabla 5B APIJhonatan Perez RamirezÎncă nu există evaluări
- Problemario de Biorreactores Cap 14Document5 paginiProblemario de Biorreactores Cap 14Nycte Nava0% (1)
- Xampler HFDocument8 paginiXampler HFAnil ReddyÎncă nu există evaluări
- Web ClientDocument117 paginiWeb ClientLim Fang JunÎncă nu există evaluări
- Elemental and Material BalanceDocument12 paginiElemental and Material BalanceGem ToledoÎncă nu există evaluări
- Analysis Synthesis and Design of Chemical Processes Third Edition T LDocument5 paginiAnalysis Synthesis and Design of Chemical Processes Third Edition T LUzair Wahid0% (1)
- Solutions Manual For Analysis Synthesis and Design of Chemical Processes 4th Edition PDFDocument14 paginiSolutions Manual For Analysis Synthesis and Design of Chemical Processes 4th Edition PDFNathalia DelgadoÎncă nu există evaluări
- Assg 4Document18 paginiAssg 4Fitria HasanahÎncă nu există evaluări
- Department of Chemical Engg. ASSIGNMENT-3, Mass Transfer IDocument3 paginiDepartment of Chemical Engg. ASSIGNMENT-3, Mass Transfer IKimberly BautistaÎncă nu există evaluări
- P5&6Document28 paginiP5&6Valeria NatashaÎncă nu există evaluări
- Semibatch UniDocument22 paginiSemibatch UniMelgi159100% (1)
- Material Balances ExercisesDocument3 paginiMaterial Balances ExercisesMigue MolinaÎncă nu există evaluări
- Ce Ic106d eDocument2 paginiCe Ic106d eredabeakÎncă nu există evaluări
- Paula Andrea Caro Baez Fluidos Ii G 2Document4 paginiPaula Andrea Caro Baez Fluidos Ii G 2PAULA ANDREA CARO BAEZÎncă nu există evaluări
- Application of Different Types of Bioreactors in BioprocessesDocument37 paginiApplication of Different Types of Bioreactors in BioprocessesKarenParadaÎncă nu există evaluări
- Vam Machine Flow DiagramDocument2 paginiVam Machine Flow DiagramDevÎncă nu există evaluări
- Soderal MNDocument7 paginiSoderal MNVictor GomezÎncă nu există evaluări
- Cap 222222Document27 paginiCap 222222Paty ChiluisaÎncă nu există evaluări
- Solución - Flash MulticomponenteDocument6 paginiSolución - Flash MulticomponenteAlberto CondeÎncă nu există evaluări
- Deshidratación y Purificación (Alcohol Absoluto)Document4 paginiDeshidratación y Purificación (Alcohol Absoluto)Jordan Venegas100% (1)
- EC1 Problemario de Equilibrio de Fases y Calculos FlashDocument2 paginiEC1 Problemario de Equilibrio de Fases y Calculos FlashRosario Zambrano CoronaÎncă nu există evaluări
- Chapter 2Document41 paginiChapter 2muhammad shahadat awanÎncă nu există evaluări
- 1984 Variables Affecting The Yield Fatty Ester From Transesterified Vegetable OilsDocument6 pagini1984 Variables Affecting The Yield Fatty Ester From Transesterified Vegetable OilsAlberto Hernández CruzÎncă nu există evaluări
- Ejercicios Cinetica Quimica Ley de VelocidadDocument6 paginiEjercicios Cinetica Quimica Ley de Velocidadjuan mosqueraÎncă nu există evaluări
- AnnieDocument6 paginiAnnieAnnie Glorina LumauigÎncă nu există evaluări
- Ejercicio Balance de Masa para SimulaciónDocument2 paginiEjercicio Balance de Masa para SimulaciónMario Albarracín0% (1)
- CH 14Document82 paginiCH 14Sadie Hnatow75% (4)
- Diffusion Effects in Enzymes Immobilized in A Porous MatrixDocument18 paginiDiffusion Effects in Enzymes Immobilized in A Porous MatrixAegee0% (1)
- Problema 12-10 TreybalDocument1 paginăProblema 12-10 TreybalMiguel Angel Lugo CarvajalÎncă nu există evaluări
- Lec 14 Mass TransferDocument36 paginiLec 14 Mass TransferWaseem abbas100% (1)
- HW 01 SolutionDocument12 paginiHW 01 Solutionmaulida rahmiÎncă nu există evaluări
- Tabla 13.1 PerryDocument5 paginiTabla 13.1 PerryKaethy VargasÎncă nu există evaluări
- Complementary Error Function Table: X Erfc (X) X Erfc (X) X Erfc (X) X Erfc (X) X Erfc (X) X Erfc (X) X Erfc (X)Document1 paginăComplementary Error Function Table: X Erfc (X) X Erfc (X) X Erfc (X) X Erfc (X) X Erfc (X) X Erfc (X) X Erfc (X)Franco Perez ReynaÎncă nu există evaluări
- Taller Parcial de ReacccionesDocument7 paginiTaller Parcial de ReacccionesAndresFelipeSotoÎncă nu există evaluări
- Tercera Asignación Control de Procesos 2021 - IIDocument7 paginiTercera Asignación Control de Procesos 2021 - IIBrayanÎncă nu există evaluări
- Solid Soluble in Tomato Products Official MethodDocument1 paginăSolid Soluble in Tomato Products Official MethodcarrietatÎncă nu există evaluări
- Tripoli University Faculty of Engineering Chemical Engineering DepartmentDocument9 paginiTripoli University Faculty of Engineering Chemical Engineering DepartmentSrewaBenshebilÎncă nu există evaluări
- Do You Like FriesDocument4 paginiDo You Like FriesRaul AparicioÎncă nu există evaluări
- Problema 5Document2 paginiProblema 5PepeArandaÎncă nu există evaluări
- Biochemical Engineering Enzyme KineticsDocument5 paginiBiochemical Engineering Enzyme KineticsLin Xian Xing33% (3)
- Enzyme KinecticsDocument22 paginiEnzyme KinecticsJohnFedericoMartinezMuñozÎncă nu există evaluări
- Chapter 6 SolutionsDocument11 paginiChapter 6 Solutionskdd1218wÎncă nu există evaluări
- 1) Condiciones de Estado Estable. 2) Conductividad Térmica y Coeficiente Convectivo Constante. 3) Flujo de Calor UnidimensionalDocument7 pagini1) Condiciones de Estado Estable. 2) Conductividad Térmica y Coeficiente Convectivo Constante. 3) Flujo de Calor UnidimensionalTamara AlbánÎncă nu există evaluări
- Evaporators 8.5-4 SolnDocument12 paginiEvaporators 8.5-4 SolnLissa HannahÎncă nu există evaluări
- TP SeminarDocument7 paginiTP SeminarKrishna Prabhu100% (1)
- 9 - Cooling Tower 2014Document20 pagini9 - Cooling Tower 2014rahemas100% (1)
- ExercisesDocument13 paginiExercisesRajpriya GuptaÎncă nu există evaluări
- TUTORIAL Mass Transfer PrincipleDocument7 paginiTUTORIAL Mass Transfer PrincipleXin-YiWoon0% (1)
- CH E Problem 3Document3 paginiCH E Problem 3Brayan AguilarÎncă nu există evaluări
- Ion Channels of Excitable Membranes: Bertil HilleDocument11 paginiIon Channels of Excitable Membranes: Bertil HilleIgnacio Martinez Garcia100% (1)
- PRQ - 2203 Clase Viernes PDFDocument8 paginiPRQ - 2203 Clase Viernes PDFJhoel SaavedraÎncă nu există evaluări
- Insulin ADocument16 paginiInsulin ACarlos JimaÎncă nu există evaluări
- QPDocument3 paginiQPgood buddyÎncă nu există evaluări
- Worked ExamplesDocument9 paginiWorked ExamplesDzung Pham0% (1)
- STD OutDocument1 paginăSTD OutTaufiq JaiÎncă nu există evaluări
- Site and Plant LayoutDocument5 paginiSite and Plant LayoutTaufiq JaiÎncă nu există evaluări
- Modul Perfect Score SBP 2009Document124 paginiModul Perfect Score SBP 2009Jia HuiÎncă nu există evaluări
- Al-Ahleia BBT (Non-Seg) PDFDocument7 paginiAl-Ahleia BBT (Non-Seg) PDFTaufiq JaiÎncă nu există evaluări
- Modul Perfect Score SBP 2009Document124 paginiModul Perfect Score SBP 2009Jia HuiÎncă nu există evaluări
- JWPN Obj-Edited 26.6.2014Document2 paginiJWPN Obj-Edited 26.6.2014Taufiq JaiÎncă nu există evaluări
- Reference 2Document7 paginiReference 2Taufiq JaiÎncă nu există evaluări
- What Is PTFEDocument2 paginiWhat Is PTFETaufiq JaiÎncă nu există evaluări
- Nsulin Kinetics, Models, and Delivery Schedules: Mones BermanDocument4 paginiNsulin Kinetics, Models, and Delivery Schedules: Mones BermanTaufiq JaiÎncă nu există evaluări
- Mini Project CHE620 - March - July 2014Document5 paginiMini Project CHE620 - March - July 2014Taufiq JaiÎncă nu există evaluări
- Lab 5: Adiabatic Production of Acetic Anhydride ObjectivesDocument1 paginăLab 5: Adiabatic Production of Acetic Anhydride ObjectivesNajwa NaqibahÎncă nu există evaluări
- LAB2Document1 paginăLAB2Taufiq JaiÎncă nu există evaluări
- Lab 1: Separation of Ammonia and WaterDocument1 paginăLab 1: Separation of Ammonia and WaterTaufiq JaiÎncă nu există evaluări
- HETP LectureDocument4 paginiHETP LectureaadipakiÎncă nu există evaluări
- Group Project - CHE604 2013Document1 paginăGroup Project - CHE604 2013Taufiq JaiÎncă nu există evaluări
- Assignment CPE624 Nov 2013Document1 paginăAssignment CPE624 Nov 2013Taufiq JaiÎncă nu există evaluări
- HETP LectureDocument4 paginiHETP LectureaadipakiÎncă nu există evaluări
- ReuseDocument5 paginiReuseTaufiq JaiÎncă nu există evaluări
- Group Project - CHE604 2013Document1 paginăGroup Project - CHE604 2013Taufiq JaiÎncă nu există evaluări
- CHE594-Mini Project AssignmentDocument1 paginăCHE594-Mini Project AssignmentTaufiq JaiÎncă nu există evaluări
- Flow Sheet NitrobenzenDocument2 paginiFlow Sheet NitrobenzenAbimata Dwi Wahyu100% (1)
- N-Propanol Draft 4 3 2 2011Document10 paginiN-Propanol Draft 4 3 2 2011Taufiq JaiÎncă nu există evaluări
- Graph N CalculationDocument10 paginiGraph N CalculationTaufiq JaiÎncă nu există evaluări
- Procedure Lab Basic WaterDocument5 paginiProcedure Lab Basic WaterTaufiq JaiÎncă nu există evaluări
- L7-Mass Balance Reactive Sytm1Document14 paginiL7-Mass Balance Reactive Sytm1Taufiq JaiÎncă nu există evaluări
- Chem NotesDocument148 paginiChem Noteskiruba devi .kÎncă nu există evaluări
- 4 Space Time SymmetriesDocument46 pagini4 Space Time Symmetriesmcb0431703Încă nu există evaluări
- Laboratory Manual: FOR Physics Laboratory - IDocument71 paginiLaboratory Manual: FOR Physics Laboratory - IAmy PetersÎncă nu există evaluări
- Cat App 003 (E)Document4 paginiCat App 003 (E)Alexander NhuanÎncă nu există evaluări
- 11 - Quadratic Forms and EllipsoidsDocument21 pagini11 - Quadratic Forms and EllipsoidsMatheus DomingosÎncă nu există evaluări
- Mathematical Modeling of Earth's Magnetic FieldDocument21 paginiMathematical Modeling of Earth's Magnetic FieldcrocomodxÎncă nu există evaluări
- Ordinary Differential EquationsDocument600 paginiOrdinary Differential EquationsJanex RebutaÎncă nu există evaluări
- Fatigue Failure Analysis - Learn EngineeringDocument3 paginiFatigue Failure Analysis - Learn Engineeringpavan317Încă nu există evaluări
- Chapter 2Document40 paginiChapter 2IntanLeeyanaÎncă nu există evaluări
- Apm2005 Book of Abstracts PDFDocument100 paginiApm2005 Book of Abstracts PDFL'enin AlephÎncă nu există evaluări
- Handout 2014bDocument74 paginiHandout 2014bChris LittleÎncă nu există evaluări
- Astm e 90Document15 paginiAstm e 90Ginneth_Millan_0Încă nu există evaluări
- Differentiation Formulas Integration FormulasDocument7 paginiDifferentiation Formulas Integration FormulasnirobÎncă nu există evaluări
- Spinodal Decomposition PDFDocument3 paginiSpinodal Decomposition PDFOmar VillanuevaÎncă nu există evaluări
- Kinetics of Catalytic Reactions: (Ch.5 Adsorption and Desorption Process)Document16 paginiKinetics of Catalytic Reactions: (Ch.5 Adsorption and Desorption Process)comeon2amÎncă nu există evaluări
- Effect of Weather Conditions On Quality of Free Space Optics LinksDocument5 paginiEffect of Weather Conditions On Quality of Free Space Optics LinksLương Xuân DẫnÎncă nu există evaluări
- CBSE Class 12 Physics Unit - 1 Syllabus: Electrostatics: Chapter - 1: Electric Charges and FieldsDocument2 paginiCBSE Class 12 Physics Unit - 1 Syllabus: Electrostatics: Chapter - 1: Electric Charges and FieldsRitesh SoniÎncă nu există evaluări
- Semyon Neskorodev+The Elbows Technique in Wing Chun PDFDocument63 paginiSemyon Neskorodev+The Elbows Technique in Wing Chun PDFAlexJKDCQCÎncă nu există evaluări
- Paper-2: Cumulative Test-1 (Ct-1)Document44 paginiPaper-2: Cumulative Test-1 (Ct-1)vikram2002Încă nu există evaluări
- D5229 D5229MDocument18 paginiD5229 D5229MEngr Khalil AkramÎncă nu există evaluări
- Physics Unit 1 6PH01 & Unit 2 6PH02 June 2009 MSDocument27 paginiPhysics Unit 1 6PH01 & Unit 2 6PH02 June 2009 MSDaniyal SiddiquiÎncă nu există evaluări
- Photolysis Study by FtirDocument8 paginiPhotolysis Study by FtirrakibhossainÎncă nu există evaluări
- Ray Theory of TransmissionDocument44 paginiRay Theory of TransmissionAkhil Arora100% (1)
- MT - Phys2 - Summer 23 - Version A - With AnswersDocument6 paginiMT - Phys2 - Summer 23 - Version A - With AnswersaamerbolookiÎncă nu există evaluări
- Strength of Materials (S.O.M.) Model Question Paper (Q.P.) SolutionDocument16 paginiStrength of Materials (S.O.M.) Model Question Paper (Q.P.) SolutionProf. P. H. Jain92% (13)
- JEE - Physics - Waves and SoundDocument16 paginiJEE - Physics - Waves and SoundSanjana KumariÎncă nu există evaluări
- TCET FE Applied Physics - I (2018-2019)Document308 paginiTCET FE Applied Physics - I (2018-2019)Kevin100% (1)
- Class 11 Physics Practice Paper 2022-23Document11 paginiClass 11 Physics Practice Paper 2022-23Curiosity SatisfiedÎncă nu există evaluări
- Safety Considerations When Lifting With Two or More CranesDocument9 paginiSafety Considerations When Lifting With Two or More CranesKyaw Kyaw AungÎncă nu există evaluări
- Optimizing Materials Cost and MechanicalDocument8 paginiOptimizing Materials Cost and MechanicalKaleem UllahÎncă nu există evaluări
- Handbook of Formulating Dermal Applications: A Definitive Practical GuideDe la EverandHandbook of Formulating Dermal Applications: A Definitive Practical GuideÎncă nu există evaluări
- Organic Chemistry for Schools: Advanced Level and Senior High SchoolDe la EverandOrganic Chemistry for Schools: Advanced Level and Senior High SchoolÎncă nu există evaluări
- The Nature of Drugs Vol. 1: History, Pharmacology, and Social ImpactDe la EverandThe Nature of Drugs Vol. 1: History, Pharmacology, and Social ImpactEvaluare: 5 din 5 stele5/5 (5)
- Periodic Tales: A Cultural History of the Elements, from Arsenic to ZincDe la EverandPeriodic Tales: A Cultural History of the Elements, from Arsenic to ZincEvaluare: 3.5 din 5 stele3.5/5 (137)
- Is That a Fact?: Frauds, Quacks, and the Real Science of Everyday LifeDe la EverandIs That a Fact?: Frauds, Quacks, and the Real Science of Everyday LifeEvaluare: 5 din 5 stele5/5 (4)
- Monkeys, Myths, and Molecules: Separating Fact from Fiction, and the Science of Everyday LifeDe la EverandMonkeys, Myths, and Molecules: Separating Fact from Fiction, and the Science of Everyday LifeEvaluare: 4 din 5 stele4/5 (1)
- AP® Chemistry Crash Course, For the 2020 Exam, Book + Online: Get a Higher Score in Less TimeDe la EverandAP® Chemistry Crash Course, For the 2020 Exam, Book + Online: Get a Higher Score in Less TimeEvaluare: 5 din 5 stele5/5 (1)
- The Periodic Table: A Very Short IntroductionDe la EverandThe Periodic Table: A Very Short IntroductionEvaluare: 4.5 din 5 stele4.5/5 (3)
- Tribology: Friction and Wear of Engineering MaterialsDe la EverandTribology: Friction and Wear of Engineering MaterialsEvaluare: 5 din 5 stele5/5 (1)
- Taste: Surprising Stories and Science About Why Food Tastes GoodDe la EverandTaste: Surprising Stories and Science About Why Food Tastes GoodEvaluare: 3 din 5 stele3/5 (20)
- AP Chemistry Flashcards, Fourth Edition: Up-to-Date Review and PracticeDe la EverandAP Chemistry Flashcards, Fourth Edition: Up-to-Date Review and PracticeÎncă nu există evaluări
- Formulating, Packaging, and Marketing of Natural Cosmetic ProductsDe la EverandFormulating, Packaging, and Marketing of Natural Cosmetic ProductsÎncă nu există evaluări
- Chemistry for Breakfast: The Amazing Science of Everyday LifeDe la EverandChemistry for Breakfast: The Amazing Science of Everyday LifeEvaluare: 4.5 din 5 stele4.5/5 (90)
- Bioplastics: A Home Inventors HandbookDe la EverandBioplastics: A Home Inventors HandbookEvaluare: 4 din 5 stele4/5 (2)
- Ingredients: A Visual Exploration of 75 Additives & 25 Food ProductsDe la EverandIngredients: A Visual Exploration of 75 Additives & 25 Food ProductsEvaluare: 4 din 5 stele4/5 (1)
- The Disappearing Spoon: And Other True Tales of Madness, Love, and the History of the World from the Periodic Table of the ElementsDe la EverandThe Disappearing Spoon: And Other True Tales of Madness, Love, and the History of the World from the Periodic Table of the ElementsEvaluare: 4 din 5 stele4/5 (146)
- Fundamentals of Chemistry: A Modern IntroductionDe la EverandFundamentals of Chemistry: A Modern IntroductionEvaluare: 5 din 5 stele5/5 (1)
- The Elements We Live By: How Iron Helps Us Breathe, Potassium Lets Us See, and Other Surprising Superpowers of the Periodic TableDe la EverandThe Elements We Live By: How Iron Helps Us Breathe, Potassium Lets Us See, and Other Surprising Superpowers of the Periodic TableEvaluare: 3.5 din 5 stele3.5/5 (22)
- Water-Based Paint Formulations, Vol. 3De la EverandWater-Based Paint Formulations, Vol. 3Evaluare: 4.5 din 5 stele4.5/5 (6)
- ICH Quality Guidelines: An Implementation GuideDe la EverandICH Quality Guidelines: An Implementation GuideAndrew TeasdaleÎncă nu există evaluări
- The Nature of Drugs Vol. 1: History, Pharmacology, and Social ImpactDe la EverandThe Nature of Drugs Vol. 1: History, Pharmacology, and Social ImpactEvaluare: 5 din 5 stele5/5 (1)