Documente Academic
Documente Profesional
Documente Cultură
Data Bel
Încărcat de
danelrey0 evaluări0% au considerat acest document util (0 voturi)
12 vizualizări13 paginiPackage provides an interface to the C++ FILEVECTOR library facilitating analysis of large (gigato tera-bytes) matrices. Primarily aimed to support genome-wide association analyses e.g. Using GenABEL,MixABEL and ProbABEL.
Descriere originală:
Drepturi de autor
© © All Rights Reserved
Formate disponibile
PDF, TXT sau citiți online pe Scribd
Partajați acest document
Partajați sau inserați document
Vi se pare util acest document?
Este necorespunzător acest conținut?
Raportați acest documentPackage provides an interface to the C++ FILEVECTOR library facilitating analysis of large (gigato tera-bytes) matrices. Primarily aimed to support genome-wide association analyses e.g. Using GenABEL,MixABEL and ProbABEL.
Drepturi de autor:
© All Rights Reserved
Formate disponibile
Descărcați ca PDF, TXT sau citiți online pe Scribd
0 evaluări0% au considerat acest document util (0 voturi)
12 vizualizări13 paginiData Bel
Încărcat de
danelreyPackage provides an interface to the C++ FILEVECTOR library facilitating analysis of large (gigato tera-bytes) matrices. Primarily aimed to support genome-wide association analyses e.g. Using GenABEL,MixABEL and ProbABEL.
Drepturi de autor:
© All Rights Reserved
Formate disponibile
Descărcați ca PDF, TXT sau citiți online pe Scribd
Sunteți pe pagina 1din 13
Package DatABEL
November 16, 2013
Type Package
Title le-based access to large matrices stored on HDD in binary format
Version 0.9-4
Date 2013-03-12
Author Yurii Aulchenko, Stepan Yakovenko, Erik Roos, Marcel Kempenaar
Maintainer Yurii Aulchenko <yurii@bionet.nsc.ru>
Depends R (>= 2.4.0), methods
Suggests GenABEL, RUnit
Description a package providing an interface to the C++ FILEVECTOR
library facilitating analysis of large (giga- to tera-bytes)
matrices; matrix storage is organized in a way that either
columns or rows are quickly accessible; primarily aimed to
support genome-wide association analyses e.g. using GenABEL,MixABEL and ProbABEL
License GPL (>= 2)
NeedsCompilation yes
Repository CRAN
Date/Publication 2013-11-16 09:10:24
R topics documented:
DatABEL-package . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 2
apply2dfo . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 2
databel . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 4
databel-class . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 4
databel2matrix . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 6
databel2text . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 7
extract_text_le_columns . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 7
1
2 apply2dfo
get_temporary_le_name . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 8
make_empty_fvf . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 8
matrix2databel . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 9
process_lm_output . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 9
text2databel . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 10
Index 13
DatABEL-package DatABEL package for fast consecutive access to large out-of-RAM
stored matrices
Description
A package interfacing FILEVECTOR C++ library for storage of and fast consecutive access to
large data matrices in out-of-RAM disk mode with regulated cache size. Columns of matrix are
accessible very quickly.
Author(s)
Yurii Aulchenko (R code), Stepan Yakovenko (R and C++ code), Andrey Chernyh (C++ code)
See Also
apply2dfo, databel2matrix, databel2text, extract_text_file_columns, matrix2databel,
text2databel, databel
apply2dfo applies a function to databel object
Description
An iterator applying a user-dened function to an object of databel-class object
Usage
apply2dfo(..., dfodata, anFUN = "lm", MAR = 2, procFUN,
outclass = "matrix", outfile, type = "DOUBLE",
transpose = TRUE)
apply2dfo 3
Arguments
dfodata databel object which is iterated over
anFUN user-dened analysis function
MAR which margin to iteracte over (default = 2, usually these are columns used to
store SNP data)
procFUN function to process the output and present that as a xed-number-of-columns
matrix or xed-length vector. Can be missing if standard functions listed below
are used. Pre-dened processors included are "process_lm_output" (can process
functions "lm", "glm", "coxph") and "process_simple_output" (process output
from "sum", "prod", "sum_not_NA" [no. non-missing obs], "sum_NA" [no.
missing obs.])
outclass output to ("matrix" or "databel")
outfile if output class is "databel", the generated object is bond to the outle
type if output class is "databel", what data tyoe to use for storage
transpose whether to transpose the output
... arguments passed to the anFUN
Value
A matrix (or databel-matrix) containing results of applying the function
Author(s)
Yurii Aulchenko
Examples
a <- matrix(rnorm(50),10,5)
rownames(a) <- paste("id",1:10,sep="")
colnames(a) <- paste("snp",1:5,sep="")
b <- as(a,"databel")
apply(a,FUN="sum",MAR=2)
apply2dfo(SNP,dfodata=b,anFUN="sum")
tA <- apply2dfo(SNP,dfodata=b,anFUN="sum",outclass="databel",outfile="tmpA")
tA
as(tA,"matrix")
apply2dfo(SNP,dfodata=b,anFUN="sum",transpose=FALSE)
tB <- apply2dfo(SNP,dfodata=b,anFUN="sum",transpose=FALSE,outclass="databel",outfile="tmpB")
tB
as(tB,"matrix")
sex <- 1*(runif(10)>.5)
trait <- rnorm(10)+sex+as(b[,2],"vector")+as(b[,2],"vector")*sex*5
apply2dfo(trait~SNP*sex,dfodata=b,anFUN="lm")
tC <- apply2dfo(trait~SNP*sex,dfodata=b,anFUN="lm",outclass="databel",outfile="tmpC")
tC
as(tC,"matrix")
apply2dfo(trait~SNP*sex,dfodata=b,anFUN="lm",transpose=FALSE)
4 databel-class
tD <- apply2dfo(trait~SNP*sex,dfodata=b,anFUN="lm",transpose=FALSE,outclass="databel",outfile="tmpD")
tD
as(tD,"matrix")
rm(tA,tB,tC,tD)
gc()
unlink("tmp*")
databel initiates databel object
Description
this is a simple wrapper for "new" function creating databel object
Usage
databel(baseobject, cachesizeMb = 64, readonly = TRUE)
Arguments
baseobject name of the le or databel-class object
cachesizeMb cache size (amount of RAM) to be used
readonly readonly ag
Author(s)
Yurii Aulchenko
databel-class Class "databel"
Description
A class interfacing FILEVECTOR C++ library class FilteredMatrix for storage of and fast consec-
utive access to large data matrices in out-of-RAM disk mode with regulated cache size. Columns
of matrix are accessible very quickly.
Objects from the Class
Objects can be created by calls of the formnew("databel", baseobject) or databel(baseobject).
FILEVECTOR data are stored using les of form BASE.fvi (index) and BASE.fvd (data). "baseob-
ject" is either the BASE name, or object of class "databel".
databel-class 5
Slots
usedRowIndex: Object of class "integer" which rows are used
usedColIndex: Object of class "integer" which columns are used
uninames: Object of class "list", containing obejcts unique.names TRUE if all dimnames are
unique; unique.colnames if column names are unique; unique.rownames if row names
are unique
backingfilename: Object of class "character" providing BASE name
cachesizeMb: Object of class "integer" size of cache to be used to access the data. If cache is
equal to the data size, the object is stored in RAM
data: Object of class "externalptr", pointer to FilteredMatrix C++ object
Methods
[ signature(x = "databel"): sub-setting object
[<- signature(x = "databel"): setting the values in the object
connect signature(object = "databel"): connects the data les to R object (calls constructor
of FilteredMatrix object, selects rows and columns)
disconnect signature(object = "databel"): disconnects the data les to R object (calls de-
structor of FilteredMatrix object, selects rows and columns)
dim signature(x = "databel"): returns dimensions of the matrix
dimnames signature(x = "databel"): returns row and column names
dimnames<- signature(x = "databel"): sets row and column names
set_dimnames<- signature(x = "databel"): sets row and column names (these could be non-
unique)
length signature(x = "databel"): returns number of elements in the matrix
save_as signature(x = "databel"): saves (a sub-set of) the object as FV-le
save_as_text signature(x = "databel"): saves (a sub-set of) the object as plain text le
backinglename signature(object = "databel"): returns BASE FILEVECTOR le name
used to store the data
cachesizeMb signature(object = "databel"): returns the cache size used
cachesizeMb<- signature(x = "databel"): sets new cache size
get_dimnames signature(object = "databel"): returns the names of rows and columns,
which may be non-unique
set_dimnames<- signature(x = "databel"): set row and column names, which may be non-
unique
setReadOnly<- signature(x = "databel"): sets ReadOnly (TRUE or FALSE) attribute to
databel object
Author(s)
Yurii Aulchenko, Stepan Yakovenko, Andrey Chernyh
6 databel2matrix
References
http://www.genabel.org/packages/DatABEL
See Also
make_empty_fvf, databel, matrix2databel
Examples
showClass("databel")
databel2matrix converts databel to matrix
Description
Converts a databel object to a regular R matrix. This is the procedure used by the "as" converting
to DatABEL objects, in which case a temporary le name is created.
Usage
databel2matrix(from, rows, cols)
Arguments
from databel matrix
rows which rows to include
cols which columns to include
Value
object of matrix class
Author(s)
Stepan Yakovenko
databel2text 7
databel2text Exports DatABEL object to a text le
Description
Exports DatABEL object to a text le
Usage
databel2text(databel, file, NAString = "NA",
row.names = TRUE, col.names = TRUE, transpose = FALSE)
Arguments
databel DatABEL object
file output le name
NAString string to replace NA with
row.names export row names if TRUE
col.names export col names if TRUE
transpose whether the matrix should be transposed
Author(s)
Stepan Yakovenko
extract_text_file_columns
extracts columns from text le
Description
Extracts a column from text le to a matrix. If in a particular le line the number of columns is less
then a column specied, returns last column!
Usage
extract_text_file_columns(file, whichcols)
Arguments
file le name
whichcols which columns to extract
Value
matrix of strings with values from that columns
8 make_empty_fvf
get_temporary_file_name
generates temporary le name
Description
function to generate temporary le name
Usage
get_temporary_file_name(path = ".", withFVext = TRUE)
Arguments
path path to directory where the temporary le will be located
withFVext whether function should check presence of *FVD and *FVI les too
make_empty_fvf makes empty levector object
Description
function to generate empty levector object (and disk les)
Usage
make_empty_fvf(name, nvariables, nobservations,
type = "DOUBLE", cachesizeMb = 64, readonly = FALSE)
Arguments
name name fo the le to be assoiated with new object
nvariables number of variables (R columns)
nobservations number of observations (R rows)
type data type of the object ("UNSIGNED_SHORT_INT", "SHORT_INT", "UN-
SIGNED_INT", "INT", "FLOAT", "DOUBLE", "CHAR", "UNSIGNED_CHAR")
cachesizeMb what cache size to use for newly generated databel object
readonly whether to open new databel in readonly mode
Value
databel object; also le is created in le system
matrix2databel 9
matrix2databel converts matrix to databel
Description
Converts regular R matrix to databel object. This is the procedure used by "as" converting to
DatABEL objects, in which case a temporary le name is created
Usage
matrix2databel(from, filename, cachesizeMb = 64,
type = "DOUBLE", readonly = FALSE)
Arguments
from R matrix
filename which FILEVECTOR BASE le name to use
cachesizeMb cache size to be used when accessing the object
type type of data to use for storage ("DOUBLE", "FLOAT", "INT", "UNSIGNED_INT",
"UNSIGNED_SHORT_INT", "SHORT_INT", "CHAR", "UNSIGNED_CHAR")
readonly whether to generate new databel in read only mode
Value
object of class databel
Author(s)
Yurii Aulchenko
process_lm_output apply2dfo-associated functions
Description
A number of functions used in conjunction with apply2dfo. Standardly supported apply2dfos
anFUN analysis functions include lm, glm, coxph, sum, prod, "sum_not_NA" (no. non-
missing obs), and "sum_NA" (no. missing obs.). Pre-dened processing functions include "pro-
cess_lm_output" (can process functions "lm", "glm", "coxph") and "process_simple_output" (pro-
cess output from "sum", "prod", "sum_not_NA", "sum_NA")
Usage
process_lm_output(lmo,verbosity=2)
10 text2databel
Arguments
lmo object returned by analysis with "lm", "glm", etc.
verbosity verbosity
See Also
apply2dfo
Examples
a <- matrix(rnorm(50),10,5)
rownames(a) <- paste("id",1:10,sep="")
colnames(a) <- paste("snp",1:5,sep="")
b <- as(a,"databel")
apply(a,FUN="sum",MAR=2)
apply2dfo(SNP,dfodata=b,anFUN="sum",procFUN="process_simple_output")
apply2dfo(SNP,dfodata=b,anFUN="sum",transpose=FALSE)
sex <- 1*(runif(10)>.5)
trait <- rnorm(10)+sex+as(b[,2],"vector")+as(b[,2],"vector")*sex*5
apply2dfo(trait~SNP*sex,dfodata=b,anFUN="lm",procFUN="process_lm_output")
text2databel converts text le to levector format
Description
The le provides the data to be converted to levector format. The le may provide the data only
(no row and column names) in which case col/row names may be left empty or provided in separate
les (in which case it is assumed that names are provided only for the imported columns/rows
see skip-options). There is an option to skip a number of rst ros and columns. The row and
column names may also be provided in the le itself, in which case one needs to tell the row/column
number providing column/row names. Unless option "R_matrix" is set to TRUE, it is asumed
that the number of columns is always the same acorss the le. If above option is provided, it
is assumed that both column and row names are provided in the le, and the rst line contains
one column less than other lines (such is the case with les produced from R using the function
write.table(...,col.names=TRUE,row.names=TRUE).
Usage
text2databel(infile, outfile, colnames, rownames,
skipcols, skiprows, transpose = FALSE,
R_matrix = FALSE, type = "DOUBLE", cachesizeMb = 64,
readonly = TRUE, naString = "NA")
text2databel 11
Arguments
infile input text le name
outfile output levector le name; if missing, it is set to inle+".levector"
colnames where are the column names stored? If missing, no column names; if integer,
this denotes the row of the input le where the column names are specied; if
character string then the string species the name of the le with column names
rownames where are the row names stored? If missing, no row names; if integer, this de-
notes the column of the input le where the row names are specied; if character
string then the string species the name of the le with row names
skipcols how many columns of the input le to skip
skiprows how many rows of the input le to skip
transpose whether the le is to be transposed
R_matrix if true, the le format is assumed to follow the format of R data matrix produced
with write.table(...,col.names=TRUE,row.names=TRUE)
type data DatABEL type to use ("DOUBLE", "FLOAT", "INT", "UNSIGNED_INT",
"UNSIGNED_SHORT_INT", "SHORT_INT", "CHAR", "UNSIGNED_CHAR")
cachesizeMb cache size for the resulting databel-class object
readonly whether the resulting databel-class object should be opened in readonly mode
naString the string used for missing data (default: NA)
Value
The converted le is stored in the le system, a databel-class object connection to the le is returned.
Author(s)
Yurii Aulchenko
Examples
cat("this is an example which you can run if you can write to the file system\n")
## Not run:
# create matrix
NC <- 5
NR <- 10
data <- matrix(rnorm(NC*NR),ncol=NC,nrow=NR)
rownames(data) <- paste("r",1:NR,sep="")
colnames(data) <- paste("c",1:NC,sep="")
data
# create text files
write.table(data,file="test_matrix_dimnames.dat",row.names=TRUE,col.names=TRUE,quote=FALSE)
write.table(data,file="test_matrix_colnames.dat",row.names=FALSE,col.names=TRUE,quote=FALSE)
write.table(data,file="test_matrix_rownames.dat",row.names=TRUE,col.names=FALSE,quote=FALSE)
write.table(data,file="test_matrix_NOnames.dat",row.names=FALSE,col.names=FALSE,quote=FALSE)
12 text2databel
write(colnames(data),file="test_matrix.colnames")
write(rownames(data),file="test_matrix.rownames")
# generate identical data
text2databel(infile="test_matrix_dimnames.dat",outfile="test_matrix_dimnames",R_matrix=TRUE)
x <- databel("test_matrix_dimnames")
data <- as(x,"matrix")
data
# convert text two filevector format
text2databel(infile="test_matrix_NOnames.dat",outfile="test_matrix_NOnames.fvf",
colnames="test_matrix.colnames",rownames="test_matrix.rownames")
x <- databel("test_matrix_NOnames.fvf")
if (!identical(data,as(x,"matrix"))) stop("not identical data")
text2databel(infile="test_matrix_NOnames.dat",outfile="test_matrix_NOnames_T.fvf",
colnames="test_matrix.colnames",rownames="test_matrix.rownames",transpose=TRUE)
x <- databel("test_matrix_NOnames_T.fvf")
if (!identical(data,t(as(x,"matrix")))) stop("not identical data")
text2databel(infile="test_matrix_rownames.dat",outfile="test_matrix_rownames.fvf",
rownames=1,colnames="test_matrix.colnames")
x <- databel("test_matrix_rownames.fvf")
if (!identical(data,as(x,"matrix"))) stop("not identical data")
text2databel(infile="test_matrix_colnames.dat",outfile="test_matrix_colnames.fvf",
colnames=1,rownames="test_matrix.rownames")
x <- databel("test_matrix_colnames.fvf")
if (!identical(data,as(x,"matrix"))) stop("not identical data")
text2databel(infile="test_matrix_dimnames.dat",outfile="test_matrix_dimnames.fvf",R_matrix=TRUE)
x <- databel("test_matrix_dimnames.fvf")
if (!identical(data,as(x,"matrix"))) stop("not identical data")
# stupid extended matrix in non-R format
newmat <- matrix(-100,ncol=NC+3,nr=NR+2)
newmat[3:(NR+2),4:(NC+3)] <- data
newmat[2,4:(NC+3)] <- paste("c",1:NC,sep="")
newmat[3:(NR+2),3] <- paste("r",1:NR,sep="")
newmat
write.table(newmat,file="test_matrix_strange.dat",col.names=FALSE,row.names=FALSE,quote=FALSE)
text2databel(infile="test_matrix_strange.dat",outfile="test_matrix_strange.fvf",
colnames=2,rownames=3)
x <- databel("test_matrix_strange.fvf")
if (!identical(data,as(x,"matrix"))) stop("not identical data")
## End(Not run)
Index
Topic classes
databel-class, 4
[,databel,ANY,ANY,ANY-method
(databel-class), 4
[,databel-method (databel-class), 4
[-methods (databel-class), 4
[<-,databel-method (databel-class), 4
apply2dfo, 2, 2, 10
backingfilename (databel-class), 4
backingfilename,databel-method
(databel-class), 4
cachesizeMb (databel-class), 4
cachesizeMb,databel-method
(databel-class), 4
cachesizeMb<- (databel-class), 4
cachesizeMb<-,databel-method
(databel-class), 4
connect (databel-class), 4
connect,databel-method (databel-class),
4
databel, 2, 4, 4, 6, 9
databel-class, 4, 4, 11
DatABEL-package, 2
databel2matrix, 2, 6
databel2text, 2, 7
dim,databel-method (databel-class), 4
dimnames,databel-method
(databel-class), 4
dimnames<-,databel-method
(databel-class), 4
disconnect (databel-class), 4
disconnect,databel-method
(databel-class), 4
extract_text_file_columns, 2, 7
get_dimnames (databel-class), 4
get_dimnames,databel-method
(databel-class), 4
get_temporary_file_name, 8
length,databel-method (databel-class), 4
make_empty_fvf, 6, 8
matrix, 6
matrix2databel, 2, 6, 9
process_lm_output, 9
process_simple_output
(process_lm_output), 9
save_as (databel-class), 4
save_as,databel-method (databel-class),
4
save_as_text (databel-class), 4
save_as_text,databel-method
(databel-class), 4
set_dimnames<- (databel-class), 4
set_dimnames<-,databel-method
(databel-class), 4
setReadOnly<- (databel-class), 4
setReadOnly<-,databel-method
(databel-class), 4
sum_NA (process_lm_output), 9
sum_not_NA (process_lm_output), 9
text2databel, 2, 10
13
S-ar putea să vă placă și
- The Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeDe la EverandThe Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeEvaluare: 4 din 5 stele4/5 (5794)
- The Little Book of Hygge: Danish Secrets to Happy LivingDe la EverandThe Little Book of Hygge: Danish Secrets to Happy LivingEvaluare: 3.5 din 5 stele3.5/5 (399)
- Shoe Dog: A Memoir by the Creator of NikeDe la EverandShoe Dog: A Memoir by the Creator of NikeEvaluare: 4.5 din 5 stele4.5/5 (537)
- Never Split the Difference: Negotiating As If Your Life Depended On ItDe la EverandNever Split the Difference: Negotiating As If Your Life Depended On ItEvaluare: 4.5 din 5 stele4.5/5 (838)
- Hidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceDe la EverandHidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceEvaluare: 4 din 5 stele4/5 (895)
- The Yellow House: A Memoir (2019 National Book Award Winner)De la EverandThe Yellow House: A Memoir (2019 National Book Award Winner)Evaluare: 4 din 5 stele4/5 (98)
- A Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryDe la EverandA Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryEvaluare: 3.5 din 5 stele3.5/5 (231)
- Grit: The Power of Passion and PerseveranceDe la EverandGrit: The Power of Passion and PerseveranceEvaluare: 4 din 5 stele4/5 (588)
- Elon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureDe la EverandElon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureEvaluare: 4.5 din 5 stele4.5/5 (474)
- On Fire: The (Burning) Case for a Green New DealDe la EverandOn Fire: The (Burning) Case for a Green New DealEvaluare: 4 din 5 stele4/5 (73)
- Team of Rivals: The Political Genius of Abraham LincolnDe la EverandTeam of Rivals: The Political Genius of Abraham LincolnEvaluare: 4.5 din 5 stele4.5/5 (234)
- The Emperor of All Maladies: A Biography of CancerDe la EverandThe Emperor of All Maladies: A Biography of CancerEvaluare: 4.5 din 5 stele4.5/5 (271)
- The Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersDe la EverandThe Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersEvaluare: 4.5 din 5 stele4.5/5 (344)
- Devil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaDe la EverandDevil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaEvaluare: 4.5 din 5 stele4.5/5 (266)
- The Unwinding: An Inner History of the New AmericaDe la EverandThe Unwinding: An Inner History of the New AmericaEvaluare: 4 din 5 stele4/5 (45)
- The World Is Flat 3.0: A Brief History of the Twenty-first CenturyDe la EverandThe World Is Flat 3.0: A Brief History of the Twenty-first CenturyEvaluare: 3.5 din 5 stele3.5/5 (2219)
- The Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreDe la EverandThe Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreEvaluare: 4 din 5 stele4/5 (1090)
- The Sympathizer: A Novel (Pulitzer Prize for Fiction)De la EverandThe Sympathizer: A Novel (Pulitzer Prize for Fiction)Evaluare: 4.5 din 5 stele4.5/5 (120)
- Her Body and Other Parties: StoriesDe la EverandHer Body and Other Parties: StoriesEvaluare: 4 din 5 stele4/5 (821)
- States of Matter PoetryDocument20 paginiStates of Matter PoetrySaqib HussainÎncă nu există evaluări
- CAP Regulation 100-1 - 08/21/2000Document16 paginiCAP Regulation 100-1 - 08/21/2000CAP History LibraryÎncă nu există evaluări
- Evolution of Information Technology and Its Impact On SocietyDocument19 paginiEvolution of Information Technology and Its Impact On SocietyKashif Naqvi100% (1)
- Limbach l2000 All Operatingandmaintenancemanual enDocument46 paginiLimbach l2000 All Operatingandmaintenancemanual enFernando MoreiraÎncă nu există evaluări
- Joining Text Tables To Replace Technical Names With Descriptions in The HANA View For Drilldown Search PDFDocument10 paginiJoining Text Tables To Replace Technical Names With Descriptions in The HANA View For Drilldown Search PDFRajaÎncă nu există evaluări
- Sap Ehp 7 For Sap Erp 6Document9 paginiSap Ehp 7 For Sap Erp 6fiestamixÎncă nu există evaluări
- Off Grid PV Systems Design GuidelinesDocument24 paginiOff Grid PV Systems Design GuidelinestantibaÎncă nu există evaluări
- Marvelous College of Technology, IncDocument7 paginiMarvelous College of Technology, Incthirsty liquidÎncă nu există evaluări
- Canon Irc3200 Parts CatalogDocument313 paginiCanon Irc3200 Parts CatalogStratis SiderisÎncă nu există evaluări
- Screw ConveyorDocument3 paginiScrew ConveyorWalter EsquivelÎncă nu există evaluări
- Chapter 14-SolutionDocument3 paginiChapter 14-Solutionmp6w9qw7t2Încă nu există evaluări
- Paragon Scientific Lubricant Certified Reference MaterialsDocument4 paginiParagon Scientific Lubricant Certified Reference MaterialsPutut WindujatiÎncă nu există evaluări
- Tanaka 2Document15 paginiTanaka 2Firani Reza ByaÎncă nu există evaluări
- Comparing Virtualization Platforms - PowerVM and VMWareDocument26 paginiComparing Virtualization Platforms - PowerVM and VMWareBart SimsonÎncă nu există evaluări
- QUY50C Electric System DiagramDocument1 paginăQUY50C Electric System DiagramMohamed SabryÎncă nu există evaluări
- b2020-Tdc-Fas-004 Fasteners r3Document2 paginib2020-Tdc-Fas-004 Fasteners r3Ramalingam PrabhakaranÎncă nu există evaluări
- Sizing a Flare Stack and Calculating Flame Distortion from Wind VelocityDocument49 paginiSizing a Flare Stack and Calculating Flame Distortion from Wind VelocityBayu AjipÎncă nu există evaluări
- FDD TDD New LayeringDocument8 paginiFDD TDD New LayeringMuhammad Arif Wibowo100% (1)
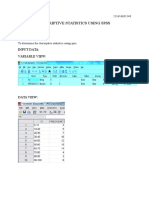
- Descriptive Statistics Using SPSS: Ex No: DateDocument5 paginiDescriptive Statistics Using SPSS: Ex No: Datepecmba11Încă nu există evaluări
- Cohesion and Adhesion in LiquidsDocument1 paginăCohesion and Adhesion in LiquidsjmlagumbayÎncă nu există evaluări
- Ddrcs Unit 4Document22 paginiDdrcs Unit 4siva shanmukhaÎncă nu există evaluări
- Gradle For Android - Sample ChapterDocument18 paginiGradle For Android - Sample ChapterPackt PublishingÎncă nu există evaluări
- Motor Insurance: Type of Policies General Regulation Rating and TariffDocument28 paginiMotor Insurance: Type of Policies General Regulation Rating and TariffArun DasÎncă nu există evaluări
- 1 Bac Global Test (U - 2 & 3)Document2 pagini1 Bac Global Test (U - 2 & 3)MouradFilaliÎncă nu există evaluări
- HISTOGRAMADocument8 paginiHISTOGRAMAGABRIELA ESTEFANIA SIGUENZA LOJAÎncă nu există evaluări
- DPWH Waterstop SpecificationDocument6 paginiDPWH Waterstop SpecificationFrancis DomingoÎncă nu există evaluări
- PCR for Keyboard EPDDocument20 paginiPCR for Keyboard EPDNurul Haerani Mohamad100% (1)
- BenQ G610HDA - V1Document47 paginiBenQ G610HDA - V1adriantxeÎncă nu există evaluări
- DR Reporting Made Easy With Report Builder 3.0Document132 paginiDR Reporting Made Easy With Report Builder 3.0robertorojasfeijoÎncă nu există evaluări
- A New and Unique Tool For Shrimp Feed Producers: Kristin Weel SundbyDocument8 paginiA New and Unique Tool For Shrimp Feed Producers: Kristin Weel SundbyrsuertoÎncă nu există evaluări