Documente Academic
Documente Profesional
Documente Cultură
Biochem Lab Exam 2 Review
Încărcat de
areddy4343100%(1)100% au considerat acest document util (1 vot)
904 vizualizări6 paginiThis document summarizes the key steps and principles of purifying recombinant green fluorescent protein (rGFP) using nickel affinity chromatography. It discusses how freezing and thawing bacterial pellets lyses cells to release rGFP, how washing removes contaminants, and how imidazole competes with the histidine tag to elute rGFP. It also reviews techniques for monitoring and quantifying purified protein concentration like UV light detection, spectrophotometry, and Bradford assays.
Descriere originală:
Biochem lab documents lab exam 2
Drepturi de autor
© © All Rights Reserved
Formate disponibile
DOCX, PDF, TXT sau citiți online pe Scribd
Partajați acest document
Partajați sau inserați document
Vi se pare util acest document?
Este necorespunzător acest conținut?
Raportați acest documentThis document summarizes the key steps and principles of purifying recombinant green fluorescent protein (rGFP) using nickel affinity chromatography. It discusses how freezing and thawing bacterial pellets lyses cells to release rGFP, how washing removes contaminants, and how imidazole competes with the histidine tag to elute rGFP. It also reviews techniques for monitoring and quantifying purified protein concentration like UV light detection, spectrophotometry, and Bradford assays.
Drepturi de autor:
© All Rights Reserved
Formate disponibile
Descărcați ca DOCX, PDF, TXT sau citiți online pe Scribd
100%(1)100% au considerat acest document util (1 vot)
904 vizualizări6 paginiBiochem Lab Exam 2 Review
Încărcat de
areddy4343This document summarizes the key steps and principles of purifying recombinant green fluorescent protein (rGFP) using nickel affinity chromatography. It discusses how freezing and thawing bacterial pellets lyses cells to release rGFP, how washing removes contaminants, and how imidazole competes with the histidine tag to elute rGFP. It also reviews techniques for monitoring and quantifying purified protein concentration like UV light detection, spectrophotometry, and Bradford assays.
Drepturi de autor:
© All Rights Reserved
Formate disponibile
Descărcați ca DOCX, PDF, TXT sau citiți online pe Scribd
Sunteți pe pagina 1din 6
1
Biochem Lab Exam 2 Review
Experiment 4- Purification of rGFP using Ni+2 Agarose
1) Underlying principle that allows for the purification of rGFP (how to purify
rGFP)?
-Exploiting the physical difference between your molecule of interest and all other
molecules.
- Chromatography separates proteins via differential interactions with both a mobile
and stationary phase.
2) Why does freezing and then thawing the bacterial pellet result in lysis?
- freezing and thawing to make crude extract
- From freezing, ice crystals form, that break open neighboring cell walls.
- Thawing helps make it more efficient because from damaged bacteria, the
cytoplasm is released in the breaking buffer. This lysozyme will lyse neighboring
bacteria and begin a chain reaction on lysing
3) Purpose of the wash step when developing the column?
- the washes pull out any contaminants and basically any proteins that arent His6-
tagged (rGFP) that dont bind to the column.
4) How are we monitoring the presence of rGFP?
-by using handheld UV light to see presencenot amount! That would be using a
fluorescent microplate reader. So the microplate reader show proteing amount and
protein activity.
If we were purifying a histidine tagged phosphatase, how would we monitor
its presence during the purification procedure?
- Monitor presence by reaction with PPP (colorimetric assay)
5) What is the underlying principle that allows for the elution of rGFP from the
Ni+2 agarose column?
- Imidazole, which is in the elution buffer, is very similar to hisitdine and competes
for the binding sites in the column. So basically, the his6 tags get released out of the
column when imidazole comes in.
6) What are the five major types of column chromatography and the underlying
scientific principles that make them useful for separating proteins?
1. Gel-filtration chromatography- separates based on molecular weight
2. Ion exchange chromatography- based on charge
a. Anion-exchange: binds negative charge
b. Cation-exchange: binds positive charge
3. Hydrophobic interaction chromatography- based on hydrophobic properties
4. Affinity chromatography- based on specific molecular binding properties.
2
- His6 tag with Ni+2 binding.
5. Partition chromatography- polarity or water solubility
Experiment 5 - Determining Total Protein Amount in rGFP Fractions
1) Based upon the design and principle of spectrophotometry, what improper
laboratory techniques could result in incorrect absorbance reading?
-Scratches, fingerprints and other residues on cuvettes can mess up readings as well
as improper orientation of the cuvette into the reader.
2) Given the conditions of an enzyme assay, be able to describe what components
of that assay should be placed in the reference cuvette to properly blank the
machine.
3) Be able to understand and correctly use the following terminology:
Wavelength- difference between two crests of a waveor any two points on a
wave.
Wavelength scan- measures the absorbance of a sample as you vary the
wavelength. Example: 260 nm/ 280 nm
Maximum wavelength- highest wavelength that displays the most absorbance.
Absorbance- amount of light absorbed by a substance at a certain wavelength.
Logarithmic measure.
Transmittance- ratio of light once it has passed through a sample to the intensity of
the light when it first entered the sample.
4) What are the differences between a spectrophotometer and a
spectrofluorometer?
- Spectrophotometer measures absorbance. Reads the amount of light the sample
absorbs at a specific wavelength.
- Spectrofluorometer measures fluorescence. Reads the amount of light the sample
generates at a specific wavelength when a different wavelength is shone on it.
5) Given then necessary data, construct and use a standard curve to identify the
amount and concentration of an unknown protein sample.
Using known protein concentration and its absorbance (ex: BSA) make standard
curve and then extrapolate for unknown.
6) Define specific activity.
In a sample, it is the ratio of activity over total amount of protein.
- RFU/ mg (of protein) make sure to convert to mg!
3
7) What are the two common procedures for quantifying protein
concentration that we used in this weeks lab?
8) What are the underlying principles of each of those procedures?
9) What are the advantages and disadvantages of each procedure?
Absorbance @ 260 nm/ 280 nm Bradford Assay
-Using spectrophotometer
-Measures absorbance max (280 nm)
caused by aa trp, tyr, and phe
- nucleic acids also absorb light at 280
so measure at 260 and use formula
-Measures absorbance of a sample with
varying wavelengths.
-standard curve to determine protein
amounts
-dye ( coomassie blue) binds primary
amines and carboxyls..Lys Arg
-the more the dye binds, the darker the
band and more absorbance
Advantages: Does not destroy your
sample, very rapid
rapid, reproducible, sensitive
Disadvantages: nucleic acids, turbidity
of solution, plus scratches on cuvette will
give false results. So you need purity!
Detergents interfere
Experiment 6- SDS/PAGE/ Commassie Blue Analysis of rGFP Fractions
1) What is the main purpose behind a SDS-PAGE gel?
To estimate the amount of a certain protein in a mixture of proteins.
- size via ladder
- purity
2) Be able to draw and correctly label a basic SDS-PAGE gel
Buffer, stacking gel, running gel, buffer
Negative electrode positive electrode
Wells, glass plates, spacer
3) What is the underlying principle behind SDS-PAGE gel?
First, denature proteins with detergent (SDS)this binds to proteins and makes
them negative.
Protein will be separated on the basis of size!...will be drawn to the positive
electrode (bottom of apparatus)
Smaller proteins move down faster, larger ones slower and on the top
Ladder helps determine molecular weight
So basically, you can tell size and how much of protein + purity is present
4) What is the purpose of SDS in the system?
Detergent, so it denatures proteins and makes them negative so they can be
separated based on size! (Smaller ones move down)
4
5) What is the purpose of the B-mercaptoethanol in the sample-loading buffer?
- helps unfold proteins by breaking disulfide bonds
- even though SDS unfolds proteins, it DOES NOT disrupt disulfide cross-links
between polypeptide chains
- usually added with SDS
- if NOT added, there will be one heavier band vs. two lighter bands
6) What is the purpose of glycerol in the sample-loading buffer?
-Basically increases the density of the sampleloading aid
-non-ionic, not reactive
-adds viscosity
7) What is the purpose of bis-acrylamide?
Polyacrylamide gels are formed by polymerization of acrylamide with the cross-
linking agent bisacrylamide. Pore size of gel can be adjusted by varying amounts of
acrylamide and bisacrylamide. Lower amounts, larger pore size. Larger the pore
size, the faster the proteins will migrate.
Gives high porosity.
Cross linking agent (helps the acrylamide polymerize)
In stacking gel- causes the samples to be concentrated into a narrow zone at the top
of the resolving gel
APS-TEMED reactioncross linkage between acrylamide
Allows separation of the SDS-coated proteins by size
8) What would happen if the chemicals listed above were left out of the system?
- glycerol: sample might not sink because the samples are not dense enough.
Keeps in the loading zone until electrophoresis starts.
- Bis-acrylamide: there will be low porosity, we will not be able to load
samples into gel. Acrylamide will not polymerize and entire thing will be
ruined. The acrylamide will polymerize into long strands and not a porous
gel.
- SDS PAGE: proteins will not denature or run through the gel because of mass
- B-mercaptoethanol: disulfide bonds will not break and any proteins that has
a bridge will oligomerize
9) Given an appropriate figure of a SDS-PAGE gel, be able to determine: mw,
%purity, yield of protein
-MW: migration distance= log mw.
-using ladder
- more mw= less migration distance
- if you are given data of MW and migrations distance (y,x). Draw trend line,
and then extrapolate data
- %purity: how much protein of interest per total amount of protein in mg= specific
activity.
5
- determine intensity of each band
- look at the boldness of color of each and estimate %
- yield: amount of protein of interest
ex: scaled, multiple by purity from actual experiment
10) Be able to construct and use a standard curve to determine the MW of a
protein.
-on y-axis log, so m=numbers are unevenly spaced out. Plot MW from ladder.
Negative slope because MW and migration distance are inversely related.
-measure bands from ladder in cm
-plot and draw standard curve
-extrapolate E3
11) Design an experiment to determine if a protein is monomeric or
oligomeric. If oligomerix, does the protein consist of only one type of subunit?
If oligomeric, are they held together by disulfide bonds?
Use B-mercaptoethanolget lighter band..without, darker
Experiment 8- SDS-PAGE/Western Blot
1) What is the main difference between staining and immunological detection?
-staining does not involve antibodies and immunological detection does
- immunological detection you use chromophore/chemi-luminescent (attached to 2
ab) and for staining, you use dye
- detection is more specific (one protein)
- staining detects all proteins
2) What are the three types of membranes discussed in class? Why is one chosen
over another?
1) Nitrocellulose: inexpensive, low tensile strength..breaks easily. Supported
nitrocellulose is a type that doesnt break as easily
2) Nylon: strong synthetic polyamide sheet, canreprobe and use multiple
reagents
3) PVDF (Polyvinylidene difluoride): high chemical resistance=high tensile
strength. High binding and retentive capacity. Expensive. Ideal for protein
sequencing and for using chemiluminescent detection system.
3) What is the purpose of incubating the membrane with BSA, gelatin, or a
mixture of dry milk?
These all are blocking reagents- saturate all the sites on the nitrocellulose that
are not already bound to the protein.
-if unoccupied sites are NOT blocked, 1 and 2 ab will bind non-specifically to
sited making it impossible to localize the protein of intrest.
-makes sure that our ab only bind where they are supposed to
6
4) What is the difference between a primary and secondary antibody?
Primary antibodies are raised against a specific antigen: ex:human anti-flu
ab
Secondary is an antibody that binds to primary antibodies. Typically labeled
with probes( chromophores) that make them useful for detection. Ex: if we
want to expose anti flu ab to goat, we would develop goat anti human flu
S-ar putea să vă placă și
- Computer Network AnswersDocument4 paginiComputer Network Answersammad ahmadÎncă nu există evaluări
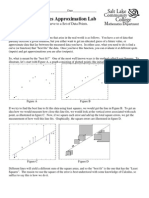
- Math 1050 Project 3 Linear Least Squares ApproximationDocument9 paginiMath 1050 Project 3 Linear Least Squares Approximationapi-233311543Încă nu există evaluări
- Acid Growth AnyakarlDocument6 paginiAcid Growth AnyakarlWillie Anne Chan UyÎncă nu există evaluări
- Lab 8 - Introduction and ProcedureDocument8 paginiLab 8 - Introduction and ProcedureFloyd Serem100% (1)
- Main 2 Master Blueprint To Language Development LatestDocument5 paginiMain 2 Master Blueprint To Language Development LatestInna Bulatova100% (1)
- MVS AP Physics C Unit 9 Lab: Hooke's Law and SHM: ObjectivesDocument2 paginiMVS AP Physics C Unit 9 Lab: Hooke's Law and SHM: ObjectivesYuyuan Luo100% (1)
- TB Chapter1Document8 paginiTB Chapter1German Calas67% (3)
- Place Value TaskDocument2 paginiPlace Value TaskKaisha MedinaÎncă nu există evaluări
- Project 2 ReportDocument9 paginiProject 2 ReportghanaÎncă nu există evaluări
- Carruthers Brute ExperienceDocument3 paginiCarruthers Brute ExperiencePatrickBrisseyÎncă nu există evaluări
- BIO 358 .01 Syllabus Spring 2021Document10 paginiBIO 358 .01 Syllabus Spring 2021Jen DongÎncă nu există evaluări
- Lab 7Document3 paginiLab 7ayaan khanÎncă nu există evaluări
- CS372 Midterm Cheat SheetDocument1 paginăCS372 Midterm Cheat SheetLim Cheng Qing100% (3)
- Lab 1 Construction of A Logic Probe: ObjectivesDocument9 paginiLab 1 Construction of A Logic Probe: ObjectivesPramote RodbonÎncă nu există evaluări
- Sydney Technical High School: Attention Mandatory Penis InspectionDocument2 paginiSydney Technical High School: Attention Mandatory Penis InspectionHarrod ThouÎncă nu există evaluări
- ColoursDocument5 paginiColoursPraveena KsÎncă nu există evaluări
- Lab 10 Radioactive Decay LawDocument5 paginiLab 10 Radioactive Decay Lawjames0% (1)
- Prepositional Phrases Adjectives and AdverbsDocument2 paginiPrepositional Phrases Adjectives and AdverbsLidisse LainezÎncă nu există evaluări
- 3Document18 pagini3Leah Nicole Erickson0% (4)
- 3DR DIY Y6 Build Manual VADocument24 pagini3DR DIY Y6 Build Manual VAFazrul100% (1)
- MB503-Practical Exam Notes VU by Muhammad KashifDocument26 paginiMB503-Practical Exam Notes VU by Muhammad KashifSagheer AhmedÎncă nu există evaluări
- 2020 - Prac 1 - SDS-PAGE and Western Blotting - BMOL3201 - 6231 - Student Notes - FINALDocument6 pagini2020 - Prac 1 - SDS-PAGE and Western Blotting - BMOL3201 - 6231 - Student Notes - FINALshaheenÎncă nu există evaluări
- DNA Restriction Enzymes Lab: Nick Milas Honors Biology May 25, 2015 Period 8Document8 paginiDNA Restriction Enzymes Lab: Nick Milas Honors Biology May 25, 2015 Period 8api-314049675Încă nu există evaluări
- Restriction MappingDocument7 paginiRestriction MappingroderickbalceÎncă nu există evaluări
- Lab Manual PDFDocument46 paginiLab Manual PDFAaron TruongÎncă nu există evaluări
- Western Blotting FINAL 2Document4 paginiWestern Blotting FINAL 2A NaÎncă nu există evaluări
- Lab Report SDS PAGEDocument8 paginiLab Report SDS PAGEHaris PapadopoulosÎncă nu există evaluări
- Lecture 22 HandoutDocument13 paginiLecture 22 HandoutPragya PandeyÎncă nu există evaluări
- Mass Spectrometry in ProteomicsDocument38 paginiMass Spectrometry in ProteomicsDawlat SalamaÎncă nu există evaluări
- The Popular Technology SDS PAGE & Western Blotting:: Principle and ApplicationDocument33 paginiThe Popular Technology SDS PAGE & Western Blotting:: Principle and ApplicationMihai GMÎncă nu există evaluări
- Principles of Separation of BiomoleculesDocument25 paginiPrinciples of Separation of BiomoleculesnikhilsathwikÎncă nu există evaluări
- Day 4 Gel ElectrophoresisDocument9 paginiDay 4 Gel Electrophoresisaguocha1Încă nu există evaluări
- Lab 2 ds180 Genotyping LabDocument7 paginiLab 2 ds180 Genotyping Labapi-342081300Încă nu există evaluări
- By The End of This Laboratory Exercise You Should Be Able ToDocument7 paginiBy The End of This Laboratory Exercise You Should Be Able TovikkyxiongÎncă nu există evaluări
- Bio Lab Assignment 2Document9 paginiBio Lab Assignment 2ElaÎncă nu există evaluări
- BSCI330 Practical Study Guide Spring 2014Document4 paginiBSCI330 Practical Study Guide Spring 2014Sahel UddinÎncă nu există evaluări
- XRF FundamentalDocument31 paginiXRF FundamentalBojan TanaskovskiÎncă nu există evaluări
- Protein Identification: Size Charge Sequence ShapeDocument34 paginiProtein Identification: Size Charge Sequence ShapeAndreea SpiridonÎncă nu există evaluări
- High Pressure/Performance Liquid Chromatography: Principle/Theory Instrumentation ApplicationDocument9 paginiHigh Pressure/Performance Liquid Chromatography: Principle/Theory Instrumentation ApplicationSubhash DhungelÎncă nu există evaluări
- Separation Techniques IDocument58 paginiSeparation Techniques ISiegfreid ArcillaÎncă nu există evaluări
- PCR Basics: Kanadi SumaprajaDocument31 paginiPCR Basics: Kanadi SumaprajaSamrichardÎncă nu există evaluări
- BIOL200 Midterm Review Session 2022WT1 - V2Document72 paginiBIOL200 Midterm Review Session 2022WT1 - V2Parveen BrarÎncă nu există evaluări
- Homologymodeling 150123025144 Conversion Gate01Document27 paginiHomologymodeling 150123025144 Conversion Gate01Jhanvi SÎncă nu există evaluări
- Experiment 5 (Lab Periods 5 and 6) Gel ElectrophoresisDocument3 paginiExperiment 5 (Lab Periods 5 and 6) Gel ElectrophoresisAli HassanÎncă nu există evaluări
- Protein Electrophoresis LabDocument8 paginiProtein Electrophoresis LabMarie St. Louis100% (1)
- Translated Version of Real Time PCRDocument8 paginiTranslated Version of Real Time PCRSonu SomanathÎncă nu există evaluări
- Real Time PCR Chemistry, Emulsion PCR and NGS PlatformDocument14 paginiReal Time PCR Chemistry, Emulsion PCR and NGS Platformnaga1975Încă nu există evaluări
- Module 2 Overview: Spring BreakDocument16 paginiModule 2 Overview: Spring BreakHaripriya SantoshÎncă nu există evaluări
- Lecture 4: Detecting Mutations and Mutation Consequences Learning GoalsDocument11 paginiLecture 4: Detecting Mutations and Mutation Consequences Learning GoalsAngelica SmithÎncă nu există evaluări
- Final PR ActsDocument26 paginiFinal PR ActsEma Smajic0% (1)
- MCAT Lab TechniquesDocument17 paginiMCAT Lab TechniquesJim Smith100% (1)
- Turbidimetry Nephelometry Model Answer 2010Document2 paginiTurbidimetry Nephelometry Model Answer 2010Bodnaru AndradaÎncă nu există evaluări
- Seen in Diagnostic Laboratories: ElectrophoresisDocument5 paginiSeen in Diagnostic Laboratories: ElectrophoresisARRIANE CYREL CAMACHOÎncă nu există evaluări
- Module IV-Screening of ClonesDocument14 paginiModule IV-Screening of ClonesAnanya SinghÎncă nu există evaluări
- Protein Mass Fingerprinting Protein Profiling Protein Assay: BIO105 / AC1Document26 paginiProtein Mass Fingerprinting Protein Profiling Protein Assay: BIO105 / AC1Christeliza Flores RaymundoÎncă nu există evaluări
- VSP UV 01 MDH Assay StudentDocument3 paginiVSP UV 01 MDH Assay StudentPhan Thanh BinhÎncă nu există evaluări
- SECT 5 SL L1-RevDocument30 paginiSECT 5 SL L1-RevUday KiranÎncă nu există evaluări
- DNA Fingerprinting Method LaboratoryDocument68 paginiDNA Fingerprinting Method Laboratorycarthagecomm28Încă nu există evaluări
- Chem Lab Report 2Document3 paginiChem Lab Report 2Maria Angela OlinanÎncă nu există evaluări
- Problem Set #2 FINALDocument5 paginiProblem Set #2 FINALgriselpaulinoÎncă nu există evaluări
- Follow The Physiology Formula Lactic AcidosisDocument59 paginiFollow The Physiology Formula Lactic AcidosisFlorin VavaÎncă nu există evaluări
- WAEC BIOLOGY SyllabusDocument78 paginiWAEC BIOLOGY SyllabusMaggieÎncă nu există evaluări
- Microbiology of Activated SludgeDocument5 paginiMicrobiology of Activated SludgeSuresh Lakshmi Narasimhan100% (1)
- Wodarg Yeadon EMA Petition Pfizer Trial FINAL 01DEC2020 en Unsigned With ExhibitsDocument43 paginiWodarg Yeadon EMA Petition Pfizer Trial FINAL 01DEC2020 en Unsigned With ExhibitsZerohedge Janitor96% (52)
- Gel Electrophoresis Lesson PlanDocument8 paginiGel Electrophoresis Lesson Planapi-215898557Încă nu există evaluări
- How Is Rhesus (RH) Typing Performed? (2019, November 10) - Retrieved March 09, 2021, From 4. HDFN More On Harmening RH FactorDocument3 paginiHow Is Rhesus (RH) Typing Performed? (2019, November 10) - Retrieved March 09, 2021, From 4. HDFN More On Harmening RH FactorValdez Francis ZaccheauÎncă nu există evaluări
- CIE Alevel Biology Mock Papers Paper 2 As Structured Questions Sample PagesDocument96 paginiCIE Alevel Biology Mock Papers Paper 2 As Structured Questions Sample PagesSalman Farsi TaharatÎncă nu există evaluări
- Biofisika NeuronDocument44 paginiBiofisika NeuronLaily KkurnyawatyÎncă nu există evaluări
- Mechanism of Action of EpinephrineDocument5 paginiMechanism of Action of EpinephrineKhalid HasanÎncă nu există evaluări
- VC Online Refresher 2017Document32 paginiVC Online Refresher 2017Angela Garcia50% (4)
- Friedland Apes CorrelationDocument1 paginăFriedland Apes Correlationapi-240829482Încă nu există evaluări
- UPDATED Annotated Cell DiagramDocument3 paginiUPDATED Annotated Cell DiagramJ pÎncă nu există evaluări
- Prospects and Challenges of Biochemistry - From The Perspective of BangladeshDocument7 paginiProspects and Challenges of Biochemistry - From The Perspective of BangladeshShimanta Easin100% (1)
- CH - 2 - Reproduction in Flowering Plants - L-1Document23 paginiCH - 2 - Reproduction in Flowering Plants - L-1yashÎncă nu există evaluări
- Bab 3 Keturunan Dan VariasiDocument13 paginiBab 3 Keturunan Dan VariasijaxsparrowÎncă nu există evaluări
- Philippine Medicinal Plants With Potential Immunomodulatory and ADocument19 paginiPhilippine Medicinal Plants With Potential Immunomodulatory and AJeaneteCaragÎncă nu există evaluări
- Microbiology v2Document1 paginăMicrobiology v2Vasili GiannoulisÎncă nu există evaluări
- W&L CatalogDocument184 paginiW&L CatalogjwwisnerÎncă nu există evaluări
- Fall 2015 Schedule of CoursesDocument15 paginiFall 2015 Schedule of CoursesThiago Antonio ZogbiÎncă nu există evaluări
- Evaluation of Food Functions and Development of Functional FoodsDocument54 paginiEvaluation of Food Functions and Development of Functional FoodsWahidah MahananiÎncă nu există evaluări
- Mammalian Oocyte Regulation Methoda and ProtocolesDocument316 paginiMammalian Oocyte Regulation Methoda and ProtocolesTlad AljazeraÎncă nu există evaluări
- Difficulty Level Analysis NEETDocument4 paginiDifficulty Level Analysis NEETVinod Kumar JainÎncă nu există evaluări
- 7.2 Gaseous Exchange in PlantsDocument17 pagini7.2 Gaseous Exchange in PlantsTheresa IzaÎncă nu există evaluări
- Wound Healing and RepairDocument54 paginiWound Healing and RepairnyangaraÎncă nu există evaluări
- Determinants of HealthDocument29 paginiDeterminants of HealthMayom MabuongÎncă nu există evaluări
- TOEFL ReadingDocument7 paginiTOEFL ReadingMaria OrlovaÎncă nu există evaluări
- Transcription and Translation VIrtual Lab Worksheet-1Document2 paginiTranscription and Translation VIrtual Lab Worksheet-1Jael Oliva EscobarÎncă nu există evaluări
- Immunological Tolerance, Pregnancy, and Preeclampsia: The Roles of Semen Microbes and The FatherDocument39 paginiImmunological Tolerance, Pregnancy, and Preeclampsia: The Roles of Semen Microbes and The FatherAnanda Yuliastri DewiÎncă nu există evaluări
- Bio2 Set BDocument22 paginiBio2 Set BAdrienaÎncă nu există evaluări
- Haemin Crystals LabDocument6 paginiHaemin Crystals LabNaiomiÎncă nu există evaluări