Documente Academic
Documente Profesional
Documente Cultură
tmp9F03 TMP
Încărcat de
FrontiersDescriere originală:
Titlu original
Drepturi de autor
Formate disponibile
Partajați acest document
Partajați sau inserați document
Vi se pare util acest document?
Este necorespunzător acest conținut?
Raportați acest documentDrepturi de autor:
Formate disponibile
tmp9F03 TMP
Încărcat de
FrontiersDrepturi de autor:
Formate disponibile
Biomed Tech 2014; 59 (s1) 2014 by Walter de Gruyter Berlin Boston. DOI 10.
1515/bmt-2014-5008
S518
Multimodal Image Segmentation of Cellular Fragmentation Using Edge
Detector and Morphological Operators
A. Khan1 , C. Weiss2 , B. Schweitzer2 , I. Hansjosten2 , R. Mikut1 and M. Reischl1
1
Institute for Applied Computer Science, Karlsruhe Institute of Technology, Germany
Institute of Toxicology and Genetics, Karlsruhe Institute of Technology, Germany
Abstract
Image processing and analysis pipelines can be adapted to solve challenging biological cell segmentation problems.
Occasionally, the image segmentation outcomes of images containing cells are unsatisfactory and insufficient if done
alone in a single channel e.g. when staining the nucleus. An efficient method using morphological operators and edge
detectors using information from an additional bright field (BF) channel is able to assign conflicting cases when segments
of nuclei belonging to a single cell are detected artificially as independent objects by nuclear staining. This method is
shown to improve the image segmentation accuracy of cell segmentation using the information from multiple channels in
comparison to only using nuclear staining.
Introduction
Currently, the focus on using image processing techniques
and methods in order to develop adequate comprehension of
biological processes has become downright attractive, both
to cell biologists and toxicologists. In multicellular organisms, different forms of cell death (such as apoptosis) are
of lasting significance for various diseases such as cancer.
Image data sets of cell populations often consist of multiple
modalities, e.g. a BF channel and a channel showing the
nuclear staining and are named according to the dye used
e.g. Hoechst (see Fig. 1). The latter channel (i.e. Hoechst)
or a channel that measures labeled green fluorescent protein (GFP) with a nuclear marker are predominantly used
in research for studying cell morphology and consequent
feature quantification (e.g. the total number of segments
detected) [1, 2]. However, if cell nuclei are fragmented due
to apoptosis, each cell consists of multiple segments (fragmented nuclei), which cannot be assigned to each other by
only using the information given in the Hoechst channel.
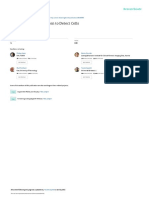
An example is given in Fig. 1. The image on the top left
shows the BF channel, the bottom left image shows the
nuclear staining (Hoechst channel) and the images on the
right show cases where stained nuclei cannot easily be recognized as fragmented nuclei belonging to one and the same
or multiple cells.
The work done related to segmentation of cells with fragmented nuclei and segmentation in the BF channel can be
seen in [3, 4, 5, 6]. However, combining multichannel information for the correct assignment of segments obtained
from a single channel needs to be explored in more detail.
Here, we propose a method to improve the segmentation
process for fragmented segments. In cases where multiple neighboring nuclear fragments occur, cell shape features
from the BF channel can be effectively used. This method
includes information from both Hoechst and BF channels
especially in cases where segments cannot be infallibly assigned. The BF channel exhibits distinctive features due to
shadows arising from membrane blebbing or cellular protrusions. The main idea is to use the segments in the BF
Figure 1: An example showing fragmented Hoechst segments in different channels : BF channel (top) and Hoechst
channel (bottom) with problematic regions highlighted in
red boxes (right side of each channel image) for both channels
channel as a measure for cellular integrity and combine this
information with the segments representing fragmented nuclei in the Hoechst channel.
Methods
The image processing pipeline implemented for this task is
shown in Fig. 2. BF channel segment information is extracted by applying a Sobel edge detector to the BF channel image. The extracted segments are further processed
sequentially using morphological operators i.e. image closing, hole-filling and image opening respectively (Fig. 3).
In the next step, binary large objects (BLOBs) detected by
Unangemeldet
Heruntergeladen am | 20.10.14 08:29
Biomed Tech 2014; 59 (s1) 2014 by Walter de Gruyter Berlin Boston. DOI 10.1515/bmt-2014-5008
S519
Figure 4: Watershed segmentation (see step 4 in Fig. 2)
based on distance map: from left to right: BLOB (highlighted in red box in Fig. 3), distance map, separated objects
ration cannot achieve the desired results due to inconsistencies caused by cellular fragmentation in both the channels.
Therefore, we use the distance map of each BLOB. As a
result all found BLOBs are within a given area range and
show convex properties (see Fig. 4).
Figure 2: Complete image processing pipeline for assign- For Hoechst segmentation, we apply a segmentation routine
described in [9, 10]. As a result we obtain i = 1, ..., n
ment of Hoechst segments to their BF counterparts
segments in the BF channel and j = 1, ..., m segments in
the Hoechst channel. The pixel positions (x/y) belonging
to the segments are gathered in the sets Bi for BF segments
and Dj for Hoechst segments. The sets D = {D1 , ..., Dm }
and B = {B1 , ..., Bn } define all found Hoechst and BF
segments respectively.
If Hoechst segments belong to the same BF segment, they
are annotated to the same class1 . Therefore, the portion pj
of positive BF segment pixels in a Hoechst segment j is
calculated and the most affecting BF segment is denoted as
nj (see Eq. 1,2 below).
T
card(Dj Bi )
pj = max
,
(1)
i
cardDj
T
card(Dj Bi )
nj = arg max
.
(2)
i
cardDj
If pj
0.1, segment Dj is removed from the Hoechst
segments and transferred to a set HOECHSTposBF segments DB. The set HOECHSTposBF contains segments in
Hoechst which are also segmented in the BF channel. All
segment sets being affected by the same BF segment are
merged:
Figure 3: BF segmentation (steps 1-2 from Fig. 2): BF (section of original BF image), 1 (edge detector), 2a (image
closing), 2b (hole filling) and 2c (image opening). Red box
highlights an example of a potentially large BLOB according to plausibility criteria and therefore would be segmented
further using step 4 in Fig. 2 i.e. watershed segmentation
(see Fig. 4)
D = {Dj |pj < 0.1},
(3)
DB = {DB1 , ..., DBn }
(4)
(5)
with
DBi =
nj =i
Dj >=0.1
Dj .
the operations 1-2 in Fig. 2 are further checked for plauEmpty sets in Eq. 4 are removed. All subsets in Eq. 3 desibility (step 3 in Fig. 2). Criteria based on irregular size
note BF-negative segments, all subsets in DB denote BFi.e. too small/big segments (approx. ranging from 200 to
positive segments. In this way segment reassignment is
1500 pixels) are implemented, thereby removing small obdone (see step 5 in Fig. 2).
jects and further segmenting big objects. A watershed seg1 If more than one BF segment affects the Hoechst segment, the most
mentation seeking for gradients in brightness is known to
perform well in cell separation [7, 8]. However, this sepa- affecting BF segment will be chosen.
Unangemeldet
Heruntergeladen am | 20.10.14 08:29
Biomed Tech 2014; 59 (s1) 2014 by Walter de Gruyter Berlin Boston. DOI 10.1515/bmt-2014-5008
S520
Results
The devised pipeline is applied to a biological dataset consisting of cis-platin treated human lung cancer cells (A549)
acquired as described earlier [11]. This dataset contains
some images of fragmented nuclei from cells that also show
some distinctive features in the BF channel. Some images
in this dataset contain 5 to 10 % of such cells. Therefore,
room to improve the segmentation accuracy is evident. The
results are shown in Fig. 5 and 6.
The goal of this processing pipeline is to annotate all segments correctly and assign only one label to all those nuclear segments that belong to one cell while giving each nucleus a unique label. All such segments belonging to each
other are assigned the same number and exact same color
in the final annotated image (a section of such an image is
shown in Fig. 5). Here, the label 77 and specific red color
tone is assigned to two segments that belong to each other. Figure 5: Output image annotated with colors and cell numTherefore, it is easier for users to instantly see which seg- bers
ments belong to each other and vice versa.
Some critical cases using different channels separately and
combined are shown in Fig. 6. Critical cases are those that
require immediate assignment but were not obtained by using only Hoechst segments. The critical cases are a combination of:
1. fragmented nuclei (as defined by manual inspection)
which however were labeled differently from each
other using Hoechst segmentation only
2. oversegmentation of the nuclei thus generating a number of Hoechst segments not corresponding to the true
number of nuclei
In Fig. 6, each column belongs to one of the several
critical cases. The first row shows critical nuclear segments
in the Hoechst channel and their corresponding appearance
in the BF channel is given in the second row. The overlaid
images are displayed in the third row and show the segments
from Hoechst in red over the grayscale BF image. The resulting annotation shows segments that belong to each other
encircled together in the final row. From Fig. 6, it is clear
that with such a method critical cases can be resolved (also
see Tab. 1).
However, the improvement in segmentation results is highly
dependent upon the number of critical cases with respect
to the total number of segments present in an image. If a
higher number of fragmented nuclei belonging to the same
cell is present, such an algorithm can highly improve the
segmentation result (see Tab. 1).
Conclusion and Outlook
This new method enables us to improve the nuclear segmentation outcome using morphological information from
another channel thereby avoiding errors in counting fragmented nuclei. The number of fragmented nuclei would
be underestimated in absence of such a method. In our
next steps, the algorithm needs to be applied to a bigger
dataset (including shading, noise etc.) to show robustness
Figure 6: Solved critical cases from given dataset (1
case per column). The first row shows critical cases in
the Hoechst channel. The second row shows the same
critical cases in BF channel. The third row represents
an overlay of the BF channel with the segments from the
Hoechst channel (shown in red color). The final row represents annotated images whereby segments belonging to
each other are encircled together (in green colored encirclements)
and performance improvement. Furthermore, implementation of automatic parameter tuning (as in big image processing pipelines) for optimization of such a method is required.
Acknowledgement
We express our gratitude to DAAD (German Academic Exchange Service) for funding this research work of A. Khan.
Unangemeldet
Heruntergeladen am | 20.10.14 08:29
Biomed Tech 2014; 59 (s1) 2014 by Walter de Gruyter Berlin Boston. DOI 10.1515/bmt-2014-5008
S521
Table 1: Results of segment reassignment and segmentation improvement. The first column shows names of images
containing critical cases. Some of these cases could be seen in Fig. 6. The second column shows the number of nuclear
segments found using the Hoechst channel only. The third column shows the number of critical cases solved by using
segmentation in the BF channel in addition to the already available Hoechst segmentation (e.g. the three critical cases
shown in Fig. 6). Here, number of cases represents the total number of critical cases in a given image and number
of segments represents the total number of segments that are involved in critical cases. The fourth column shows the
total number of segments (belonging to critical cases) that are now assigned correctly using this methodology e.g., in
column 1 and row 4 of Fig. 6 the two lower segments are now correctly assigned to the same cell shown within green
encirclements. However, not in all critical cases the improved algorithm could correctly assign the segments as manual
inspection revealed (listed in Tab. 1 as incorrect reassignment). The last column shows the improvement in segmentation
results using this method. Here, the first sub-column is assigned to the % improvement in segmentation results using the
total number of correct segments with respect to the total number of segments detected originally in a given image. This
percentage is simply calculated using: 100 (1 - (total Hoechst segments without using BF channel - difference of correct
and incorrect number of segments after reassignment)) / total Hoechst segments without using BF channel. The second
sub-column is dedicated to segmentation success with respect to the number of critical cases solved. This percentage is
simply calculated as: 100 (segments reassigned correctly - segments reassigned incorrectly) / total number of reassigned
segments per image. NOTE: no. and w.r.t are the abbreviations for number and with respect to respectively.
Image name
E1P2
H1P3
H3P4
H8P4
H9P1
H9P4
total segments
from Hoechst
266
82
40
88
104
138
critical cases
no. of cases no. of segments
11
24
1
2
7
16
4
9
7
17
5
11
References
segments reassigned
correct
incorrect
9
4
1
0
8
1
5
0
7
3
5
1
segmentation correction (%)
w.r.t image w.r.t critical cases
1.88
69.2
1.22
100
17.5
88.89
5.80
100
3.85
70
2.90
83.3
Blagosklonny, K. Blomgren, C. Borner, et al., Guidelines for the use and interpretation of assays for monitoring cell death in higher eukaryotes, Cell Death &
Differentiation, vol. 16, no. 8, pp. 10931107, 2009.
[1] B. Neumann, T. Walter, J.-K. Hrich, J. Bulkescher,
H. Erfle, C. Conrad, P. Rogers, I. Poser, M. Held,
U. Liebel, et al., Phenotypic profiling of the human
genome by time-lapse microscopy reveals cell division [7] S. Beucher and F. Meyer, Mathematical Morphology in
Image Processing, ch. The morphological approach to
genes, Nature, vol. 464, no. 7289, pp. 721727, 2010.
segmentation: The watershed transformation., pp. 433
[2] J. Stegmaier, J. C. Otte, A. Kobitski, A. Bartschat,
481. 1993.
A. Garcia, G. U. Nienhaus, U. Straehle, and R. Mikut,
[8] A. Pinidiyaarachchi and C. Whlby, Seeded waFast segmentation of stained nuclei in terabyte-scale,
tersheds for combined segmentation and tracking of
time resolved 3d microscopy image stacks, PLOS
cells, in Image Analysis and ProcessingICIAP 2005,
ONE, vol. 9, no. 2, p. e90036, 2014.
pp. 336343, Springer, 2005.
[3] F. Buggenthin, C. Marr, M. Schwarzfischer, P. S.
[9] A. Khan, M. Reischl, B. Schweitzer, C. Weiss, and
Hoppe, O. Hilsenbeck, T. Schroeder, and F. J. Theis,
R. Mikut, Feedback-driven design of normalization
An automatic method for robust and fast cell detectechniques for biological images using fuzzy formulation in bright field images from high-throughput mition of a priori knowledge, Studies in Computational
croscopy, BMC bioinformatics, vol. 14, no. 1, p. 297,
Intelligence, vol. 445, pp. 167178, 2013.
2013.
[10] A. Khan, R. Mikut, B. Schweitzer, C. Weiss, and
[4] Z. Bacs, R. B. Everson, and J. F. Eliason, The DNA
M. Reischl, Automatic tuning of image segmentation
of annexin v-binding apoptotic cells is highly fragparameters by means of fuzzy feature evaluation, Admented, Cancer Research, vol. 60, no. 16, pp. 4623
vances in Intelligent Systems and Computing, vol. 190,
4628, 2000.
no. 5, pp. 459467, 2013.
[5] S. Henery, T. George, B. Hall, D. Basiji, W. Ortyn,
[11] J. Donauer, I. Schreck, U. Liebel, and C. Weiss, Role
and P. Morrissey, Quantitative image based apoptotic
and interaction of p53, bax and the stress-activated proindex measurement using multispectral imaging flow
tein kinases p38 and jnk in benzo(a)pyrene-diolepoxide
cytometry: A comparison with standard photometric
induced apoptosis in human colon carcinoma cells,
methods, Apoptosis, vol. 13, no. 8, pp. 10541063,
Archives of Toxicology, vol. 86(2), pp. 329337, 2012.
2008.
[6] L. Galluzzi, S. A. Aaronson, J. Abrams, E. S. Alnemri,
D. W. Andrews, E. H. Baehrecke, N. G. Bazan, M. V.
Unangemeldet
Heruntergeladen am | 20.10.14 08:29
S-ar putea să vă placă și
- Image Segmentation Based On Improved UnetDocument7 paginiImage Segmentation Based On Improved UnetZAHRA FASKAÎncă nu există evaluări
- Standard and Super-Resolution Bioimaging Data Analysis: A PrimerDe la EverandStandard and Super-Resolution Bioimaging Data Analysis: A PrimerÎncă nu există evaluări
- Multi-Cell Cooperation For Future Wireless Systems: A. Silva, R. Holakouei and A. GameiroDocument23 paginiMulti-Cell Cooperation For Future Wireless Systems: A. Silva, R. Holakouei and A. Gameirojesus1843Încă nu există evaluări
- Automatic Vessel Extraction With Combined Bottom Hat and Match FilterDocument5 paginiAutomatic Vessel Extraction With Combined Bottom Hat and Match FilterBahareh Bozorg ChamiÎncă nu există evaluări
- International Journal of Engineering Research and Development (IJERD)Document10 paginiInternational Journal of Engineering Research and Development (IJERD)IJERDÎncă nu există evaluări
- Error Concealment Techniques For Video Transmission Over Error-Prone Channels: A SurveyDocument12 paginiError Concealment Techniques For Video Transmission Over Error-Prone Channels: A SurveyWalid El-ShafaiÎncă nu există evaluări
- Robust Fuzzy C-Mean Algorithm For Segmentation and Analysis of Cytological ImagesDocument3 paginiRobust Fuzzy C-Mean Algorithm For Segmentation and Analysis of Cytological ImagesWARSE JournalsÎncă nu există evaluări
- Recognition and Counting of Wbcs Using Wavelet TransformDocument5 paginiRecognition and Counting of Wbcs Using Wavelet TransformInternational Journal of Application or Innovation in Engineering & ManagementÎncă nu există evaluări
- Article Conférence ADRAR 2017Document5 paginiArticle Conférence ADRAR 2017Benomar AmineÎncă nu există evaluări
- Treatment Planning PaperDocument8 paginiTreatment Planning Paperapi-598487146Încă nu există evaluări
- Methods For Monitoring Cellular Motion and FunctionDocument4 paginiMethods For Monitoring Cellular Motion and FunctionDiogo MoreiraÎncă nu există evaluări
- Simultaneous Fusion, Compression, and Encryption of Multiple ImagesDocument7 paginiSimultaneous Fusion, Compression, and Encryption of Multiple ImagessrisairampolyÎncă nu există evaluări
- Paper Title (Use Style: Paper Title) : Subtitle As Needed (Paper Subtitle)Document6 paginiPaper Title (Use Style: Paper Title) : Subtitle As Needed (Paper Subtitle)Ahasan UllaÎncă nu există evaluări
- Currently Available Maxillofacial CBCT EquipmentDocument4 paginiCurrently Available Maxillofacial CBCT EquipmentzilniÎncă nu există evaluări
- DSA Image Registration Based On Multiscale Gabor Filters and Mutual InformationDocument6 paginiDSA Image Registration Based On Multiscale Gabor Filters and Mutual InformationerdoganaaaÎncă nu există evaluări
- A Block-Based Inter-Band Lossless Hyperspectral Image CompressorDocument10 paginiA Block-Based Inter-Band Lossless Hyperspectral Image Compressorrnagu1969Încă nu există evaluări
- Software Defect Identification Using MacDocument10 paginiSoftware Defect Identification Using Macwmarasigan2610Încă nu există evaluări
- Abd0957 SMDocument61 paginiAbd0957 SMShivaprakash Jagalur MuttÎncă nu există evaluări
- Vera 2019 J. Phys. Conf. Ser. 1160 012001Document8 paginiVera 2019 J. Phys. Conf. Ser. 1160 012001veramig1396Încă nu există evaluări
- v1s1 4Document8 paginiv1s1 4Editorijset IjsetÎncă nu există evaluări
- Novel Feedback Bit Allocation Methods For Multi-Cell Joint Processing SystemsDocument7 paginiNovel Feedback Bit Allocation Methods For Multi-Cell Joint Processing SystemsBala KumarÎncă nu există evaluări
- Multi-modality Cardiac Imaging: Processing and AnalysisDe la EverandMulti-modality Cardiac Imaging: Processing and AnalysisPatrick ClarysseÎncă nu există evaluări
- Compusoft, 3 (8), 1053-1058 PDFDocument6 paginiCompusoft, 3 (8), 1053-1058 PDFIjact EditorÎncă nu există evaluări
- 205 Automatic Segmentation of HeadDocument11 pagini205 Automatic Segmentation of HeadHead RadiotherapyÎncă nu există evaluări
- Shahad Alward Treatment Planning PaperDocument9 paginiShahad Alward Treatment Planning Paperapi-691667702Încă nu există evaluări
- Statistics: 1-Completely Randomized DesignDocument20 paginiStatistics: 1-Completely Randomized Designمصطفى الصومالÎncă nu există evaluări
- An Automatic Diagnostic System For CT Liver Image ClassificationDocument12 paginiAn Automatic Diagnostic System For CT Liver Image ClassificationShankar GaneshÎncă nu există evaluări
- Dekker Biophysj 20151Document9 paginiDekker Biophysj 20151Divya MattaÎncă nu există evaluări
- Model Based and Iconic Multimodal Registration To Merge Gated Cardiac PET, CT and MR SequencesDocument4 paginiModel Based and Iconic Multimodal Registration To Merge Gated Cardiac PET, CT and MR SequencesAnaAAAMÎncă nu există evaluări
- Study of Denoising Algorithm For MR Image Based On Contourlet TransformDocument4 paginiStudy of Denoising Algorithm For MR Image Based On Contourlet TransformSukhmander SinghÎncă nu există evaluări
- Dos 523 ProjectDocument13 paginiDos 523 Projectapi-375157618Încă nu există evaluări
- 8.nagabhushana - Nagabhushanrm@Document5 pagini8.nagabhushana - Nagabhushanrm@Ademar Alves TrindadeÎncă nu există evaluări
- Writing SampleDocument14 paginiWriting SampleYiming JingÎncă nu există evaluări
- Wavelet Based Noise Reduction in CT-Images Using Correlation AnalysisDocument20 paginiWavelet Based Noise Reduction in CT-Images Using Correlation Analysisprashant jadhavÎncă nu există evaluări
- 3-D Phantom To Simulate Cerebral Blood Flow and Metabolic Images For PetDocument5 pagini3-D Phantom To Simulate Cerebral Blood Flow and Metabolic Images For PetDaniel TervalánÎncă nu există evaluări
- FFT in Imaging ProcessDocument5 paginiFFT in Imaging ProcessJose AriasÎncă nu există evaluări
- Radiation Oncology: Investigation of The Usability of Conebeam CT Data Sets For Dose CalculationDocument13 paginiRadiation Oncology: Investigation of The Usability of Conebeam CT Data Sets For Dose CalculationCarolina CorreiaÎncă nu există evaluări
- Draft: Tof/Rgb Sensor Fusion For 3-D EndosDocument12 paginiDraft: Tof/Rgb Sensor Fusion For 3-D EndosSauravÎncă nu există evaluări
- Quantitative Cone-Beam CT Imaging in Radiotherapy Parallel Computation and Comprehensive Evaluation On The TrueBeam SystemDocument8 paginiQuantitative Cone-Beam CT Imaging in Radiotherapy Parallel Computation and Comprehensive Evaluation On The TrueBeam SystemKe LuÎncă nu există evaluări
- Automatic Lung Segmentation in CT Images Using Watershed TransformDocument4 paginiAutomatic Lung Segmentation in CT Images Using Watershed TransformPrakash SinghÎncă nu există evaluări
- Comment On "Observing The Average Trajectories of Single Photons in A Two-Slit Interferometer"Document8 paginiComment On "Observing The Average Trajectories of Single Photons in A Two-Slit Interferometer"Turing DanielaÎncă nu există evaluări
- Chapter 1 IntroductionDocument52 paginiChapter 1 IntroductionSiva sai VenkateshÎncă nu există evaluări
- Pendahuluan Id enDocument5 paginiPendahuluan Id enSri SujiatiÎncă nu există evaluări
- Supporting Online Material For: Generation of Spatial Patterns Through Cell Polarity SwitchingDocument18 paginiSupporting Online Material For: Generation of Spatial Patterns Through Cell Polarity SwitchingmetrofrogÎncă nu există evaluări
- Individual Dosimetry System For Targeted Alpha TheDocument8 paginiIndividual Dosimetry System For Targeted Alpha TheMauricio García CamachoÎncă nu există evaluări
- An Introduction To Microarray AnalysisDocument25 paginiAn Introduction To Microarray AnalysisSaurav Sarkar100% (1)
- Extraction of Coronary Vessel Structures in Low Quality X-Ray Angiogram Images Cemal Köse, Cevat İkibaşDocument4 paginiExtraction of Coronary Vessel Structures in Low Quality X-Ray Angiogram Images Cemal Köse, Cevat İkibaşvanessagirondaÎncă nu există evaluări
- Research Paper 1Document11 paginiResearch Paper 1khizrullahkhanÎncă nu există evaluări
- Removal of Blocking Artifacts of DCT Transform by Classied Space-Frequency FilteringDocument5 paginiRemoval of Blocking Artifacts of DCT Transform by Classied Space-Frequency FilteringTaz SahilÎncă nu există evaluări
- Identifying Tuberculosis Type in CtsDocument8 paginiIdentifying Tuberculosis Type in Ctswork gradsÎncă nu există evaluări
- You Should Use Regression To Detect Cells: October 2015Document9 paginiYou Should Use Regression To Detect Cells: October 2015Vint PineÎncă nu există evaluări
- Medical Imaging ThesisDocument2 paginiMedical Imaging ThesismiraclesureshÎncă nu există evaluări
- Video Coding SchemeDocument12 paginiVideo Coding SchemezfzmkaÎncă nu există evaluări
- Xmatrix White PaperDocument7 paginiXmatrix White PaperRicardo Vivancos DelgadoÎncă nu există evaluări
- MRI For Attenuation Correction in PET: Methods and ChallengesDocument15 paginiMRI For Attenuation Correction in PET: Methods and ChallengesOmarah AbdalqaderÎncă nu există evaluări
- Laser Scanning CytometryPrinciplesDocument26 paginiLaser Scanning CytometryPrinciplesSandra HernándezÎncă nu există evaluări
- (2021J) 09350305 NDocument11 pagini(2021J) 09350305 Nmohammed reyasÎncă nu există evaluări
- A Simulation-Based Assessment of The Revised NEMA NU-2 70-cm Long Test Phantom For PETDocument5 paginiA Simulation-Based Assessment of The Revised NEMA NU-2 70-cm Long Test Phantom For PETMarcelo ToledoÎncă nu există evaluări
- Image Deringing Using Quadtree Based Block-Shift FilteringDocument4 paginiImage Deringing Using Quadtree Based Block-Shift FilteringSridhar SomappaÎncă nu există evaluări
- Modelling Optimization and Control of Biomedical SystemsDe la EverandModelling Optimization and Control of Biomedical SystemsÎncă nu există evaluări
- tmpF178 TMPDocument15 paginitmpF178 TMPFrontiersÎncă nu există evaluări
- tmp3CAB TMPDocument16 paginitmp3CAB TMPFrontiersÎncă nu există evaluări
- tmpEFCC TMPDocument6 paginitmpEFCC TMPFrontiersÎncă nu există evaluări
- tmp80F6 TMPDocument24 paginitmp80F6 TMPFrontiersÎncă nu există evaluări
- tmp6F0E TMPDocument12 paginitmp6F0E TMPFrontiersÎncă nu există evaluări
- tmpCE8C TMPDocument19 paginitmpCE8C TMPFrontiersÎncă nu există evaluări
- tmpE3C0 TMPDocument17 paginitmpE3C0 TMPFrontiersÎncă nu există evaluări
- Tmpa077 TMPDocument15 paginiTmpa077 TMPFrontiersÎncă nu există evaluări
- Tmp1a96 TMPDocument80 paginiTmp1a96 TMPFrontiersÎncă nu există evaluări
- tmpE7E9 TMPDocument14 paginitmpE7E9 TMPFrontiersÎncă nu există evaluări
- tmpF3B5 TMPDocument15 paginitmpF3B5 TMPFrontiersÎncă nu există evaluări
- tmpB1BE TMPDocument9 paginitmpB1BE TMPFrontiersÎncă nu există evaluări
- tmpFFE0 TMPDocument6 paginitmpFFE0 TMPFrontiersÎncă nu există evaluări
- tmpF407 TMPDocument17 paginitmpF407 TMPFrontiersÎncă nu există evaluări
- tmp37B8 TMPDocument9 paginitmp37B8 TMPFrontiersÎncă nu există evaluări
- tmp6382 TMPDocument8 paginitmp6382 TMPFrontiersÎncă nu există evaluări
- tmp72FE TMPDocument8 paginitmp72FE TMPFrontiersÎncă nu există evaluări
- tmp998 TMPDocument9 paginitmp998 TMPFrontiersÎncă nu există evaluări
- tmp8B94 TMPDocument9 paginitmp8B94 TMPFrontiersÎncă nu există evaluări
- tmpC0A TMPDocument9 paginitmpC0A TMPFrontiersÎncă nu există evaluări
- tmpD1FE TMPDocument6 paginitmpD1FE TMPFrontiersÎncă nu există evaluări
- tmpA0D TMPDocument9 paginitmpA0D TMPFrontiersÎncă nu există evaluări
- tmp9D75 TMPDocument9 paginitmp9D75 TMPFrontiersÎncă nu există evaluări
- tmp60EF TMPDocument20 paginitmp60EF TMPFrontiersÎncă nu există evaluări
- tmp4B57 TMPDocument9 paginitmp4B57 TMPFrontiersÎncă nu există evaluări
- tmpC30A TMPDocument10 paginitmpC30A TMPFrontiersÎncă nu există evaluări
- Tmp75a7 TMPDocument8 paginiTmp75a7 TMPFrontiersÎncă nu există evaluări
- tmp3656 TMPDocument14 paginitmp3656 TMPFrontiersÎncă nu există evaluări
- tmp27C1 TMPDocument5 paginitmp27C1 TMPFrontiersÎncă nu există evaluări
- tmp2F3F TMPDocument10 paginitmp2F3F TMPFrontiersÎncă nu există evaluări
- Welcome To All: Nursing StaffDocument67 paginiWelcome To All: Nursing StaffMukesh Choudhary JatÎncă nu există evaluări
- Cardiac Cycle: DR Rakesh JainDocument97 paginiCardiac Cycle: DR Rakesh JainKemoy FrancisÎncă nu există evaluări
- Evaluation of Anti Stress Effects of Nardostachys Jatamansi DC Root Extract On Clinical Patients A Psycological EstimationDocument8 paginiEvaluation of Anti Stress Effects of Nardostachys Jatamansi DC Root Extract On Clinical Patients A Psycological EstimationESSENCE - International Journal for Environmental Rehabilitation and ConservaionÎncă nu există evaluări
- Dystolic Dysfunction Ppt. SalmanDocument42 paginiDystolic Dysfunction Ppt. SalmanMustajab MujtabaÎncă nu există evaluări
- Food Industry TrainingDocument18 paginiFood Industry TrainingSumit KumarÎncă nu există evaluări
- Robert Hooke (1665) - Who Named The Biological Unit Cell Cell Theory 1839 - byDocument11 paginiRobert Hooke (1665) - Who Named The Biological Unit Cell Cell Theory 1839 - byKamlesh RatnamÎncă nu există evaluări
- Is Fast Food The New Tobacco PDFDocument6 paginiIs Fast Food The New Tobacco PDFCustom Writing ServicesÎncă nu există evaluări
- Assessing The Ear and HearingDocument32 paginiAssessing The Ear and HearingArlyn Mendenilla0% (1)
- Transient Tachypnea of The Newborn (TTN)Document6 paginiTransient Tachypnea of The Newborn (TTN)Wivan Havilian DjohanÎncă nu există evaluări
- FNCP (Open Drainage) For Soft BoundDocument3 paginiFNCP (Open Drainage) For Soft BoundSean Maghinay BanicoÎncă nu există evaluări
- Sop Icu H & FW 12725 28.04.2018Document20 paginiSop Icu H & FW 12725 28.04.2018shah007zaadÎncă nu există evaluări
- Areport 08Document247 paginiAreport 08Mithilesh JhaÎncă nu există evaluări
- 5.RTRI Principle and Performance 8may2019Document34 pagini5.RTRI Principle and Performance 8may2019ABCDeÎncă nu există evaluări
- Lifebuoy 120324115017 Phpapp02Document38 paginiLifebuoy 120324115017 Phpapp02Binush ThampanÎncă nu există evaluări
- Excretion and Homeostasis.Document70 paginiExcretion and Homeostasis.philomenanjuguna72Încă nu există evaluări
- MedicalerrorDocument38 paginiMedicalerrorchanda jabeenÎncă nu există evaluări
- Kendriya Vidyalaya Kirandul: Investigatory Project of BiologyDocument10 paginiKendriya Vidyalaya Kirandul: Investigatory Project of BiologyAachal GayreÎncă nu există evaluări
- Diagnostic Stethoscope Project ReportDocument81 paginiDiagnostic Stethoscope Project ReportBhanu Pratap Reddy100% (1)
- Molecular Biology of BacteriaDocument57 paginiMolecular Biology of BacteriaSean ArifinÎncă nu există evaluări
- Challenges of Catholic Doctors in The Changing World - 15th AFCMA Congress 2012Document218 paginiChallenges of Catholic Doctors in The Changing World - 15th AFCMA Congress 2012Komsos - AG et al.Încă nu există evaluări
- Icru Report 62Document62 paginiIcru Report 62Luis Ramirez100% (1)
- A Drug Study On PrednisoneDocument5 paginiA Drug Study On PrednisonePrincess Alane MorenoÎncă nu există evaluări
- CENE Course Outline On Psychiatric NursingDocument3 paginiCENE Course Outline On Psychiatric NursingJohn Ryan BuenaventuraÎncă nu există evaluări
- Fidelis Drug List 2018Document80 paginiFidelis Drug List 2018Annie AnnaÎncă nu există evaluări
- Bilirubin Metabolism Jaundice PPP 9Document27 paginiBilirubin Metabolism Jaundice PPP 9Regina Marsha Xaveria100% (1)
- Congenital Insensitivity To Pain With Anhidrosis (CIPA) : A Case ReportDocument5 paginiCongenital Insensitivity To Pain With Anhidrosis (CIPA) : A Case ReportAhmad DiazÎncă nu există evaluări
- Magic of The Minimum Dose PDFDocument221 paginiMagic of The Minimum Dose PDFminunat100% (1)
- Atrial Fibrillation PathophysiologyDocument2 paginiAtrial Fibrillation PathophysiologyanabbabarÎncă nu există evaluări
- Samar Deb Easy and Interesting ADocument846 paginiSamar Deb Easy and Interesting ACharlieÎncă nu există evaluări
- Women and Mental HealthDocument74 paginiWomen and Mental Healthmerin sunilÎncă nu există evaluări