Documente Academic
Documente Profesional
Documente Cultură
Ieee
Încărcat de
HelloprojectDrepturi de autor
Formate disponibile
Partajați acest document
Partajați sau inserați document
Vi se pare util acest document?
Este necorespunzător acest conținut?
Raportați acest documentDrepturi de autor:
Formate disponibile
Ieee
Încărcat de
HelloprojectDrepturi de autor:
Formate disponibile
2017 4th International Conference on Signal Processing, Communications and Networking (ICSCN -2017), March 16 – 18, 2017,
Chennai, INDIA
Medical Image Processing to Detect Breast
Cancer - A Cognitive-Based Investigation
1 2
D. Saranyaraj M. Manikandan
Research Scholar, Department of Electronics Associate Professor, Department of
Engineering, Anna University, MIT Campus, Electronics Engineering, Anna University,
Chennai, TamilNadu. MIT Campus, Chennai, TamilNadu.
Abstract- This paper expounds the cognitive Image image processing is that the data of the image
processing techniques involved in medical images. developed would not trade despite the fact that it's
There are several varieties of proliferative cancer that reconstructed a number of occasions protecting the
are being well-known by means of the researchers,
originality of the image [1]. The medicals images
which has now hit the tally to a hundred. Each and
have got to be built with accuracy, displaying
every cancer is unique of its sort to be famous
alongside the signs. This paper specializes in the more
instantly for the surgeon to research, automatic and
than a few image processing algorithms involved in correct segmentation of elements of interest,
diagnosing the breast melanoma which is perilous Quantitative size of image points and an
cancer prompted in Women worldwide. The statistics interpretation of the measurements[2]
for this gain knowledge of is taken from the
International agency of research on cancer- World There are distinct forms of cancer which range
Health Organization and American cancer society. The up to 100 and extra. The most damaging and
benchmark of the present and prior study is shown to customary cancer are a Breast, Lung, Prostate [3].
enhance the long run findings. This Paper specializes in one such perilous cancer
that is Breast melanoma. The additional sections of
Keywords- Cancer Detection; Breast cancer; Early
Detection of Cancer; Image Processing.
this paper extract the current research achievements
by the researchers on more than a few image
I. INTRODUCTION processing systems concerned in detecting breast
cancer from the mammogram pictures. This survey
Medical Image evaluation is important in the paper will inspire the extra investigations in the
area of remedy. With a tremendous quantity of medical image processing.
patient information, numerous scientific functions
require a big knowledge to be processed. This paper
makes a specialty of one such analysis which is on
melanoma. It's identified that 21.7 million number of
latest cancer circumstances are anticipated to be
identified in 2030. And, by 2030, 13 million
melanoma deaths are predicted. This file was
released with the aid of Global Cancer Facts &
Figures, 3rd Edition, produced by the American
Cancer Society incorporation with IARC, on
February 4, 20151. There are lots of invasive
methods in diagnosing melanoma like
Mammography, Histopathology, biopsy, X-Rays,
CT-Scan, etc., these approaches both emit radiation
which is hazardous or exerts pressure. Fig. 1. Leading Sites of New Cancer Cases and Deaths – 2016
Estimates [1].
The image processing strategies have been
developed for a correct evaluation on detecting more
than a few melanoma. Informed technicians are
required to diagnose the scan results along with the
mutation of melanoma cells2, 3, which can also be
minimized with the image processing algorithms.
Clearly, the important thing benefit of the Digital
978-1-5090-4740-6/17/$31.00 ©2017 IEEE
2017 4th International Conference on Signal Processing, Communications and Networking (ICSCN -2017), March 16 – 18, 2017,
Chennai, INDIA
II. INSIGHTS OF BREAST CANCER
Table 2. Breast Cancer Incidence -Women in India (2012-2015)
Melanoma is the most dangerous sickness
[3]
which happens due to the harm of the tissues in any
part of the physique. Breast cancer affects one in
eight females of their lifetime. Clinical research
concentrating on breast melanoma shouldn't be new
and its roots return into the sixteenth century. Due to
the lack of conversation and advancement in the
scientific area, this ailment regarded as one of the
deadly illnesses of all of the occasions. The tissue is
categorized as benign and malignant tissues, where
benign remains restricted to the web page of its
origin and does no longer unfold to different
materials of the physique with confined harm & is
non-cancerous. Whereas, malignant first grows
slowly without signs. This stage is called the latent
stage. The tumor later grows speedily. The cancer
2015 Breast Cancer Rate of incidence -
cells go past adjoining tissue and enter the blood and
Estimated number of Breast cancer
India
lymph. Once this happens, they migrate to many
different sites in the body the place the cancer cells Incidence (All ages) 200000 155863
proceed to divide. The lifetime danger of incidence 144937
150000
or Mortality from melanoma refers back to the 100000
10926
danger a character has, over the path of his or her 50000 0 0 0 0 0 0 Male
lifetime, of being diagnosed with or death from 0
Female
melanoma.
Both sex
The following table 1, table 2, Figure 2&3 gives
the incidence rate and the mortality rate of the
Year of Incidence
Breast cancer in Women only, classifying into two
age groups, below 65 years and above 65 years. The
demographic trade within the populace of the cancer Figure 2. The statistical representation of the Incidence rate in
incidence is given from which the change within the Breast Cancer in India(2012-2015) [3]
cancer incidence between the year 2012 and 2015
[3]. 2015 Breast Cancer Rate of Mortality -
Estimated number of Breast
cancer Mortality (All ages)
India
Table 1. Breast Cancer Incidence -Women in India (2012-2015)
[3] 70218
80000 75957
60000 5739
40000 0 0 0
20000
0 Male
Female
Both sex
Year of Mortality
Figure 3. The statistical representation of the Incidence rate in
Breast Cancer in India(2012-2015) [3]
III. ANOMALY DETECTION OF
MAMMOGRAPHY USING MULTIPLE-
INSTANCE LEARNING
978-1-5090-4740-6/17/$31.00 ©2017 IEEE
2017 4th International Conference on Signal Processing, Communications and Networking (ICSCN -2017), March 16 – 18, 2017,
Chennai, INDIA
Gwenol´e Quellec et al.[4], proposed CAD based technique. The computed areas beneath receiver
detection and diagnosis for mammography running characteristic curves monotonically
examinations. A method for partitioning the breast expanded from 0.666 ± zero.029 to 0.730 ± 0.027as
relatively and automatically into areas was the time-lag among the “previous” (three to at least
proposed. Within the proposed framework, breasts one) and “cutting-edge” examinations decreases. the
are first sectioned into regions. The normal and very best adjusted odds ratios were 5.63, 7.43Forty
abnormal areas derived from facets from the 3, and eleven.1 for the three “earlier” (3 to 1 units of
laceration and textural characteristics. When an examinations, respectively. this is taught verified an
abnormality is detected, the regions of prognosis are optimistic association among the chance rankings
highlighted. Two approaches are weighed up to generated by means of a bilateral mammographic
delineate this incongruity detector. In the second characteristic trade targeted risk mannequin and a
state of affairs, the neighborhood and world faulty growing fashion of the near-term hazard for having
cells are trained at the same time, without handbook mammography-detected breast melanoma.
segmentations, using more than a few Multiple-
Table 3
Instance Learning MIL algorithms. The Digital Baseline characteristics of positive (cancer) and negative (cancer-
Database for Screening Mammography (DDSM) free) cases [5] of 335 cases. Source: Maxine Tan et al., [5]
dataset exhibit that the 2d strategy, which is handiest
weakly supervised, signifies the primary method,
although it's strongly-supervised. This implies that
fault detection can be informed on significant
medical report archives, discarding the need for
handbook segmentation.
IV. EARLY PREDICTION OF BREAST
CANCER ASSOCIATED WITH
MAMMOGRAPHIC IMAGE FEATURES.
Maxine Tan et al. [5], designed a computerized
model to predict the early detection of breast cancer
to be diagnosed by comparing the previous
mammogram image characteristics from a series of
Multiple-Instance Learning FFDM images which is
shown in Table 3. The dataset incorporates a
sequence of 4 chronological FFDM results of 335
women. The remaining conclusion from each and
each progression (“present”) and the 3-brand new
“previous” examinations have been got. All
“previous” examinations were interpreted as bad V. STACKED SPARSE AUTOENCODER
within the route of the ordinary scientific image (SSAE) ANALYSIS ON NUCLEI
studying, whilst inside the “present” examinations DETECTION FROM THE
159 cancers were observed and confirmed. 176 HISTOPATHOLOGY IMAGES OF BREAST
epochs remaining melanoma-unfastened. From CANCER
every picture, the writer firstly estimated 158
mammographic densities, structural similarity, and Computerized nuclear detection is a crucial step for
texture located photo features. Absolutely the some of PC assisted pathology related photo
subtraction fee among the left and right breasts was analysis algorithms corresponding to for
selected to represent each and each characteristic. computerized grading of breast cancer tissue
The author then built three support vector machine specimens. J. Xu et al. [6] proposed a stacked Sparse
(SVM) situated danger objects that have been Autoencoder (SSAE), an example of a deep
knowledgeable and established using a leave-one- mastering method, is supplied for efficient nuclei
case-out situated bypass-validation gadget. The real detection on excessive-selection histopathological
points utilized in each SVM model were decided on pictures of breast melanoma. The SSAE learns
utilizing a nested stepwise regression evaluation excessive-degree elements from absolutely pixel
978-1-5090-4740-6/17/$31.00 ©2017 IEEE
2017 4th International Conference on Signal Processing, Communications and Networking (ICSCN -2017), March 16 – 18, 2017,
Chennai, INDIA
intensities on its own as a manner to decide breast parenchyma 3) Extraction of
distinguishing factors of nuclei. A sliding window texture elements like suggest, average deviation,
operation is applied to every image to be able to entropy, kurtosis and so forth,
constitute image patches through excessive-degree geometric features like area, perimeter, L:S, ENC,
factors acquired thru the Autoencoder, which may (Elliptical normalized circumference)
be then therefore fed to a classifier which wavelet located factors , just so signatures can also
categorizes each image patch as nuclear or be assigned to identity and class of benign and
nonnuclear. The author considered 500 malignant masses. Fourteen sufferers’
histopathological images with 2200*2200 dimension mammograms had been processed. features of
with 3500 segmented nuclei characteristics based on six sufferers were extracted that have hundreds. It is
the reality. Hence SSAE will show the result located that a lot of these points help to categorize
pertaining to 84:49% and aveP as 78:83%. Hence benign and malignant lots. If many parameters are
the author concluded that SSAE is better performed extracted for an extra number of sufferers who have
than the other trending nuclear detection techniques. plenty, a mannequin may also be developed to
Thus those nuclei are detected using various categorize benign and malignant instances which
approaches based on categories given in Table 4. facilitate the healthcare professional to further
examine the instances for early detection of breast
Table 4
Different Nuclei Detection Approaches with their Categories melanoma in the shorter time. In future, we might
advocate ways to extract points of plenty and
Microcalcifications and extra classify the breast
mass and Microcalcifications to identify benign or
malignant sufferers for early detection of breast
melanoma.
VII. BREAST TUMOR DETECTION IN
DIGITAL MAMMOGRAPHY BASED ON
EXTREME LEARNING MACHINE
Zhiqiong Wang et.al proposed a breast tumor
detection algorithm in digital mammography on
ELM. Primarily, image preprocessing is performed
to cancel the noise for which the median filter was
used and image enhancement was performed
basically to improve the contrast of the digital
mammography in knowledge preprocessing. The
morphological and region of interest method was
sequentially implemented for the CC view of the
breast images for wavelet modulus maxima
canceling the unwanted parts and for selecting the
required region of interest for the classification
purpose. After which textural elements (5) and
VI. DETECTION OF BREAST CANCER IN morphological features (5) are extracted. Based on
AN EARLY STAGE USING VIRTUAL the features and textures extracted the ELM
INSTRUMENTATION [35] algorithm is performed to detect the breast
malignant tissue. Comparing breast tumor detection
Spandana Paramkusham et.al. Proposed a unique set based on help Vector Machines (SVM), with breast
of rules for early detection of tumor detection established on ELM, not only does
breast cancer utilizing image processing techniques. ELM have a greater classification accuracy than
Novel algorithms are carried out for 1) SVM, however, it also has a generally multiplied
Mass location extraction to get exact form of the training pace.
mass 2) Superposition of boundary of mass on
mammogram helps docs to view the
boundary effortlessly as mass location overlaps with
978-1-5090-4740-6/17/$31.00 ©2017 IEEE
2017 4th International Conference on Signal Processing, Communications and Networking (ICSCN -2017), March 16 – 18, 2017,
Chennai, INDIA
Table 5
Analysis of Various Algorithms with its Advantages
VIII. CONCLUSION cut down the mortality price by additionally
detecting in an early stage of prevalence.
This paper elaborates distinctively about more than
a few State-of-artwork issues with quite a lot of REFERENCE
algorithms and approaches involving detection of [1]. D.A.Mahalle , Prof. V.N.Dhage, " Machine Vision based
Breast melanoma. Various Machine learning Approach for Cordial Image Processing", International
Algorithms like Backpropagation, Stacked Engineering Journal For Research & Development, Vol 2,
Autoencoder, multiple instance learning, canny Issue 3.
[2]. Dougherty Geoff. 15 June 2011. Medical Image
edge detection had been discussed which has both processing Techniques and Applications. Springer New
merits and demerits. It is inferred that identifying York. DOI 10.1007/978-1-4419-9779-1_1.
the overlap of cells, abnormalities detection, and [3]. American Cancer Society. Global Cancer Facts & Figures
accuracy rate and image enhancement can be more 3rd Edition. Atlanta: American Cancer Society; 2015.
[4]. G. Quellec, M. Lamard, M. Cozic, G. Coatrieux and G.
efficient when artificial neural networks algorithms Cazuguel, "Multiple-Instance Learning for Anomaly
are implemented. Also using the virtual Detection in Digital Mammography," in IEEE
Transactions on Medical Imaging, vol. 35, no. 7, pp. 1604-
instrumentation the detection of cancer can be 1614, July 2016
performed at an early stage of occurrence. This [5]. M. Tan, B. Zheng, J. K. Leader and D. Gur, "Association
overall review will facilitate in bettering the long Between Changes in Mammographic Image Features and
Risk for Near-Term Breast Cancer Development," in IEEE
run research problems. Established on this survey
Transactions on Medical Imaging, vol. 35, no. 7, pp. 1719-
Detection of Breast melanoma is primary so as to 1728, July 2016hgdfbn
978-1-5090-4740-6/17/$31.00 ©2017 IEEE
2017 4th International Conference on Signal Processing, Communications and Networking (ICSCN -2017), March 16 – 18, 2017,
Chennai, INDIA
[6]. J. Xu et al., "Stacked Sparse Autoencoder (SSAE) for [22]. F. Xing, H. Su, J. Neltner, and L. Yang, “Automatic ki- 67
Nuclei Detection on Breast Cancer Histopathology counting using robust cell detection and online dictionary
Images," in IEEE Transactions on Medical Imaging, vol. learning,” Biomedical Engineering, IEEE Transactions on,
35, no. 1, pp. 119-130, Jan. 2016. vol. 61, no. 3, pp. 859–870, March 2014.
[7]. H. Chang, J. Han, A. Borowsky, L. Loss, J.W. Gray, P.T. [23]. C. Jung and C. Kim, “Segmenting clustered nuclei using
Spellman, and B. Parvin, “Invariant delineation of nuclear hminima transform-based marker extraction and contour
architecture in glioblastoma multiforme for clinical and parameterization,” Biomedical Engineering, IEEE
molecular association,” Medical Imaging, IEEE Transactions on, vol. 57, no. 10, pp. 2600–2604, Oct 2010.
Transactions on, vol. 32, no. 4, pp. 670–682, April 2013. [24]. Y. Yan, X. Zhou, M. Shah, and S.T.C. Wong, “Automatic
[8]. H. Irshad, S. Jalali, L. Roux, D. Racoceanu, L. J. Hwee, G. segmentation of high-throughput rnai fluorescent cellular
Le Naour, and F. Capron, “Automated mitosis detection images,” Information Technology in Biomedicine, IEEE
using texture, sift features and hmax biologically inspired Transactions on, vol. 12, no. 1, pp. 109–117, Jan 2008.
approach,” Journal of Pathology informatics, vol. 4, no. [25]. S. Ali and A. Madabhushi, “An integrated region-,
Suppl, 2013. boundary-, shape-based active contour for multiple object
[9]. S. D. Cataldo, E. Ficarra, and E. Acquaviva, A.and Macii, overlap resolution in histological imagery.,” IEEE Trans
“Automated segmentation of tissue images for Med Imaging, vol. 31, no. 7, pp. 1448–1460, July 2012.
computerized ihc analysis,” Computer Methods and [26]. C. Yan, A. Li, B. Zhang, W. Ding, Q. Luo, and H. Gong,
Programs in Biomedicine, vol. 100, no. 1, pp. 1 – 15, “Automated and accurate detection of soma location and
2010. surface morphology in large-scale 3d neuron images,” PLoS
[10]. M. Oberlaender, V. J. Dercksen, R. Egger, R. Gensel, B. ONE, vol. 8, no. 4, pp. e62579, 04 2013.
Sakmann, and H. Hege, “Automated three-dimensional [27]. H. Esmaeilsabzali, K. Sakaki, N. Dechev, R. Burke, and E.J.
detection and counting of neuron somata,” Journal of Park, “Machine vision-based localization of nucleic and
Neuroscience Methods, vol. 180, no. 1, pp. 147 – 160, 2009. cytoplasmic injection sites on low-contrast adherent cells,”
[11]. B. Nielsen, F. Albregtsen, and H. E. Danielsen, “Automatic Medical & Biological Engineering & Computing, vol. 50,
segmentation of cell nuclei in feulgen-stained histological no. 1, pp. 11–21, 2012.
sections of prostate cancer and quantitative evaluation of [28]. P. Filipczuk, T. Fevens, A Krzyzak, and R. Monczak,
segmentation results,” Cytometry Part A, vol. 81, no. 7, pp. “Computer-aided breast cancer diagnosis based on the
588–601, 2012. analysis of cytological images of fine needle biopsies,”
[12]. A.C. Ruifrok and D. A. Johnston, “Quantification of Medical Imaging, IEEE Transactions on, vol. 32, no. 12,
histochemical staining by color deconvolution.,” Analytical pp. 2169–2178, Dec 2013.
and quantitative cytology and histology/the International [29]. A.Basavanhally, S. Ganesan, S. Agner, J. P. Monaco, M. D.
Academy of Cytology [and] American Society of Cytology, Feldman, J. E. Tomaszewski, G. Bhanot, and A.
vol. 23, no. 4, pp. 291–299, 2001. Madabhushi, “Computerized image-based detection and
[13]. J. P. Vink, M. B. Van Leeuwen, C. H. M. Van Deurzen, and grading of lymphocytic infiltration in her2+ breast cancer
G. De Haan, “Efficient nucleus detector in histopathology histopathology,” Biomedical Engineering, IEEE
images,” Journal of microscopy, vol. 249, no. 2, pp. 124– Transactions on, vol. 57, no. 3, pp. 642–653, 2010.
135, 2013. [30]. E. Bernardis and S. X. Yu, “Pop out many small structures
[14]. S. Petushi, F. U. Garcia, M. M. Haber, C. Katsinis, and A. from a very large microscopic image,” Medical Image
Tozeren, “Large-scale computations on histology images Analysis, vol. 15, no. 5, pp. 690 – 707, 2011, Special Issue
reveal grade-differentiating parameters for breast cancer,” on the 2010 Conference on Medical Image Computing and
BMC Medical Imaging, vol. 6, no. 1, pp. 14, 2006. Computer- Assisted Intervention.
[15]. H. Fatakdawala, J. Xu, A. Basavanhally, G. Bhanot, S. [31]. G. M. Faustino, M. Gattass, C.J.P. de Lucena, P. B.
Ganesan, M. Feldman, J.E. Tomaszewski, and A. Campos, and S. K. Rehen, “A graph-mining algorithm for
Madabhushi, “Expectation maximization-driven geodesic automatic detection and counting of embryonic stem cells in
active contour with overlap resolution (emagacor): fluorescence microscopy images,” Integrated Computer-
Application to lymphocyte segmentation on breast cancer Aided Engineering, vol. 18, no. 1, pp. 91–106, 2011.
histopathology,” Biomedical Engineering, IEEE [32]. G. Li, T. Liu, J. Nie, L. Guo, J. Malicki, A. Mara, S. A.
Transactions on, vol. 57, no. 7, pp. 1676–1689, 2010. Holley, W. Xia, and S.T.C. Wong, “Detection of blob
[16]. J. Byun, M. R. Verardo, B. Sumengen, G. P Lewis, B. S. objects in microscopic zebrafish images based on gradient
Manjunath, and S. K. Fisher, “Automated tool for the vector diffusion,” Cytometry Part A, vol. 71, no. 10, pp.
detection of cell nuclei in digital microscopic images: 835–845, 2007.
application to retinal images,” Mol Vis, vol. 12, pp. 949– [33]. T. Liu, G. Li, J. Nie, A. Tarokh, X. Zhou, L. Guo, J.
960, 2006. Malicki, W. Xia, and S.T.C. Wong, “An automated method
[17]. Y. Al-Kofahi, W. Lassoued, W. Lee, and B. Roysam, for cell detection in zebrafish,” Neuroinformatics, vol. 6, no.
“Improved automatic detection and segmentation of cell 1, pp. 5–21, 2008.
nuclei in histopathology images,” Biomedical Engineering, [34]. A.QCruz-Roa, J. Arevalo Ovalle, A. Madabhushi, and F.
IEEE Transactions on, vol. 57, no. 4, pp. 841–852, April Gonzalez Osorio, “A deep learning architecture for image
2010. representation, visual interpretability, and automated basal-
[18]. B. Parvin, Q. Yang, J. Han, H. Chang, B. Rydberg, and cell carcinoma cancer detection,” in MICCAI 2013,
M.H. Barcellos-Hoff, “Iterative voting for inference of Kensaku Mori, Ichiro Sakuma, Yoshinobu Sato, Christian
structural saliency and characterization of subcellular Barillot, and Nassir Navab, Eds., vol. 8150 of Lecture Notes
events,” Image Processing, IEEE Transactions on, vol. 16, in Computer Science, pp. 403– 410. Springer Berlin
no. 3, pp. 615–623, March 2007. Heidelberg, 2013.
[19]. X. Qi, Y. Xing, D.J. Foran, and L. Yang, “Robust [35]. S. Paramkusham, K. M. M. Rao and B. V. V. S. N.
segmentation of overlapping cells in histopathology Prabhakar Rao, "Early stage detection of breast cancer using
specimens using parallel seed detection and repulsive level novel image processing techniques, Matlab and Labview
set,” Biomedical Engineering, IEEE Transactions on, vol. implementation," 2013 15th International Conference on
59, no. 3, pp. 754–765, March 2012. Advanced Computing Technologies (ICACT), Rajampet,
[20]. O. Schmitt and M. Hasse, “Radial symmetries based 2013, pp. 1-5.
decomposition of cell clusters in binary and gray level [36]. Zhiqiong Wang, GeYub, YanKang, YingjieZhao , QixunQu
images,” Pattern Recognition, vol. 41, no. 6, pp. 1905 – "Breast tumor detection in digital mammography based on
1923, 2008. extreme learning machine", Neurocomputing128(2014)175–
[21]. H. Wang, F. Xing, H. Su, A. Stromberg, and L. Yang, 184.
“Novel image markers for non-small cell lung cancer
classification and survival prediction,” BMC Bioinformatics,
vol. 15, no. 1, pp. 310, 2014.
978-1-5090-4740-6/17/$31.00 ©2017 IEEE
S-ar putea să vă placă și
- PostDocument1 paginăPostHelloprojectÎncă nu există evaluări
- Suresh Front PageDocument12 paginiSuresh Front PageHelloprojectÎncă nu există evaluări
- SystemDocument1 paginăSystemHelloprojectÎncă nu există evaluări
- Java-Project Title 2021Document6 paginiJava-Project Title 2021HelloprojectÎncă nu există evaluări
- Level-0: Airline Agency Customer Maintanence Provide ServiceDocument2 paginiLevel-0: Airline Agency Customer Maintanence Provide ServiceHelloprojectÎncă nu există evaluări
- Resumes Anja I Kanna MRDocument2 paginiResumes Anja I Kanna MRHelloprojectÎncă nu există evaluări
- Project Abstract ON A Gaussian Derivative Based Version of Jpeg For Image Compressionand DecompressionDocument2 paginiProject Abstract ON A Gaussian Derivative Based Version of Jpeg For Image Compressionand DecompressionHelloprojectÎncă nu există evaluări
- Mathematics ReportDocument9 paginiMathematics ReportHelloprojectÎncă nu există evaluări
- Sdkhfuishfiosdhfsdfs DFKGDJGDFKHGDF DFGKDFGDF DFKGJDFKGDocument1 paginăSdkhfuishfiosdhfsdfs DFKGDJGDFKHGDF DFGKDFGDF DFKGJDFKGHelloprojectÎncă nu există evaluări
- Sis Sonn PDFDocument4 paginiSis Sonn PDFHelloprojectÎncă nu există evaluări
- Light-Weight Multi-Document Summarization Based On Two-Pass Re-RankingDocument7 paginiLight-Weight Multi-Document Summarization Based On Two-Pass Re-RankingHelloprojectÎncă nu există evaluări
- Syn-Data Structure TutorDocument1 paginăSyn-Data Structure TutorHelloprojectÎncă nu există evaluări
- Mib Final LeafDocument12 paginiMib Final LeafHelloprojectÎncă nu există evaluări
- Project Abstract ON A Gaussian Derivative Based Version of Jpeg For Image Compressionand DecompressionDocument2 paginiProject Abstract ON A Gaussian Derivative Based Version of Jpeg For Image Compressionand DecompressionHelloprojectÎncă nu există evaluări
- License PremiumDocument1 paginăLicense PremiumNguyen Viet Trung (FPL HCMK13.3)100% (1)
- Padal Left and Right TurningDocument1 paginăPadal Left and Right TurningHelloprojectÎncă nu există evaluări
- Project Abstract ON A Gaussian Derivative Based Version of Jpeg For Image Compressionand DecompressionDocument2 paginiProject Abstract ON A Gaussian Derivative Based Version of Jpeg For Image Compressionand DecompressionHelloprojectÎncă nu există evaluări
- Nithya Prasath New Resume 2018-HelloDocument3 paginiNithya Prasath New Resume 2018-HelloHelloprojectÎncă nu există evaluări
- NishaDocument36 paginiNishaHelloprojectÎncă nu există evaluări
- Front MIBDocument8 paginiFront MIBHelloprojectÎncă nu există evaluări
- Design and Fabrication of Eco-Friendly Aqua SilencerDocument7 paginiDesign and Fabrication of Eco-Friendly Aqua SilencerIJSTEÎncă nu există evaluări
- Table of ContentsDocument1 paginăTable of ContentsHelloprojectÎncă nu există evaluări
- Data Entry - Agreement FormDocument3 paginiData Entry - Agreement FormHelloprojectÎncă nu există evaluări
- Padal Left and Right TurningDocument1 paginăPadal Left and Right TurningHelloprojectÎncă nu există evaluări
- Nithya Prasath New Resume 2018-HelloDocument3 paginiNithya Prasath New Resume 2018-HelloHelloprojectÎncă nu există evaluări
- Letter 3Document1 paginăLetter 3HelloprojectÎncă nu există evaluări
- RfererDocument1 paginăRfererHelloprojectÎncă nu există evaluări
- New Microsoft Office Word DocumentDocument1 paginăNew Microsoft Office Word DocumentHelloprojectÎncă nu există evaluări
- ComparativeDocument60 paginiComparativeAdeelÎncă nu există evaluări
- Table of ContentsDocument1 paginăTable of ContentsHelloprojectÎncă nu există evaluări
- The Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeDe la EverandThe Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeEvaluare: 4 din 5 stele4/5 (5794)
- Shoe Dog: A Memoir by the Creator of NikeDe la EverandShoe Dog: A Memoir by the Creator of NikeEvaluare: 4.5 din 5 stele4.5/5 (537)
- Hidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceDe la EverandHidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceEvaluare: 4 din 5 stele4/5 (895)
- The Yellow House: A Memoir (2019 National Book Award Winner)De la EverandThe Yellow House: A Memoir (2019 National Book Award Winner)Evaluare: 4 din 5 stele4/5 (98)
- The Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersDe la EverandThe Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersEvaluare: 4.5 din 5 stele4.5/5 (344)
- The Little Book of Hygge: Danish Secrets to Happy LivingDe la EverandThe Little Book of Hygge: Danish Secrets to Happy LivingEvaluare: 3.5 din 5 stele3.5/5 (399)
- Grit: The Power of Passion and PerseveranceDe la EverandGrit: The Power of Passion and PerseveranceEvaluare: 4 din 5 stele4/5 (588)
- The Emperor of All Maladies: A Biography of CancerDe la EverandThe Emperor of All Maladies: A Biography of CancerEvaluare: 4.5 din 5 stele4.5/5 (271)
- Devil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaDe la EverandDevil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaEvaluare: 4.5 din 5 stele4.5/5 (266)
- Never Split the Difference: Negotiating As If Your Life Depended On ItDe la EverandNever Split the Difference: Negotiating As If Your Life Depended On ItEvaluare: 4.5 din 5 stele4.5/5 (838)
- A Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryDe la EverandA Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryEvaluare: 3.5 din 5 stele3.5/5 (231)
- On Fire: The (Burning) Case for a Green New DealDe la EverandOn Fire: The (Burning) Case for a Green New DealEvaluare: 4 din 5 stele4/5 (73)
- Elon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureDe la EverandElon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureEvaluare: 4.5 din 5 stele4.5/5 (474)
- Team of Rivals: The Political Genius of Abraham LincolnDe la EverandTeam of Rivals: The Political Genius of Abraham LincolnEvaluare: 4.5 din 5 stele4.5/5 (234)
- The Unwinding: An Inner History of the New AmericaDe la EverandThe Unwinding: An Inner History of the New AmericaEvaluare: 4 din 5 stele4/5 (45)
- The World Is Flat 3.0: A Brief History of the Twenty-first CenturyDe la EverandThe World Is Flat 3.0: A Brief History of the Twenty-first CenturyEvaluare: 3.5 din 5 stele3.5/5 (2259)
- The Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreDe la EverandThe Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreEvaluare: 4 din 5 stele4/5 (1090)
- The Sympathizer: A Novel (Pulitzer Prize for Fiction)De la EverandThe Sympathizer: A Novel (Pulitzer Prize for Fiction)Evaluare: 4.5 din 5 stele4.5/5 (121)
- Her Body and Other Parties: StoriesDe la EverandHer Body and Other Parties: StoriesEvaluare: 4 din 5 stele4/5 (821)
- GS Oral SDSDocument108 paginiGS Oral SDSrobhopp67% (3)
- Surgical Specialty Oral Q's (Somewhat Translated)Document3 paginiSurgical Specialty Oral Q's (Somewhat Translated)dÎncă nu există evaluări
- Dermatology Questions and AnsDocument147 paginiDermatology Questions and AnsEl FaroukÎncă nu există evaluări
- Breslow. English Name. 2016Document2 paginiBreslow. English Name. 2016Anca-Raluca PascuÎncă nu există evaluări
- Case Series of Rare Breast Diseases and Their Unusual PresentationDocument36 paginiCase Series of Rare Breast Diseases and Their Unusual Presentationdodo1Încă nu există evaluări
- Uworld - SURGERYDocument55 paginiUworld - SURGERYNikxyÎncă nu există evaluări
- Skin DisordersDocument202 paginiSkin DisordersMj Briones100% (1)
- Benign Neoplasms and HyperplasiasDocument19 paginiBenign Neoplasms and HyperplasiasimperiouxxÎncă nu există evaluări
- Benign Skin TumoursDocument7 paginiBenign Skin TumoursMan LorÎncă nu există evaluări
- Ocular Melanoma: The Eye Signs and SymptomsDocument2 paginiOcular Melanoma: The Eye Signs and SymptomsPutra TridiyogaÎncă nu există evaluări
- Local Flap Reconstruction of Large Scalp DefectsDocument5 paginiLocal Flap Reconstruction of Large Scalp DefectsPalwasha MalikÎncă nu există evaluări
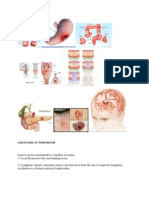
- Cancer Mode of TransmissionDocument4 paginiCancer Mode of Transmission9chand3100% (1)
- Fitzpatrick Skin TypeDocument4 paginiFitzpatrick Skin Typenoor hidayahÎncă nu există evaluări
- Specialty Certificate in Medical Oncology Sample QuestionsDocument27 paginiSpecialty Certificate in Medical Oncology Sample Questionsvimal raj100% (1)
- Citologia VeterinariaDocument19 paginiCitologia VeterinariaJuan Manuel ValenciaÎncă nu există evaluări
- 2.malignant Tumours of Epithelial Cell Origin-2Document86 pagini2.malignant Tumours of Epithelial Cell Origin-2Dinesh Kr. YadavÎncă nu există evaluări
- Ddg.13546.Doc BenerDocument25 paginiDdg.13546.Doc Benernailatul fadhilaÎncă nu există evaluări
- PosterDocument45 paginiPosterRoxana Alexandra BogosÎncă nu există evaluări
- Epidemiology of Ocular Tumors 2013Document230 paginiEpidemiology of Ocular Tumors 2013Ranny LaidasuriÎncă nu există evaluări
- 989104Document188 pagini989104diyan110Încă nu există evaluări
- Epidemiology of Cutaneous Melanoma in Germany and Worldwide: Claus Garbe Andreas BlumDocument11 paginiEpidemiology of Cutaneous Melanoma in Germany and Worldwide: Claus Garbe Andreas BlumGabriela AgrigÎncă nu există evaluări
- A Natural Anti-Cancer ProtocolDocument12 paginiA Natural Anti-Cancer Protocolstraw man100% (1)
- Skin and Endo 2022Document10 paginiSkin and Endo 2022Sure NavyasriÎncă nu există evaluări
- DUB Brochure English DB10 2012 ODocument16 paginiDUB Brochure English DB10 2012 OJing CruzÎncă nu există evaluări
- SURGERY CASES MADE EASY MedicineDocument106 paginiSURGERY CASES MADE EASY MedicineShamal100% (3)
- Cadth Issues in Emerging: Health TechnologiesDocument20 paginiCadth Issues in Emerging: Health TechnologiesKrishna SrihariÎncă nu există evaluări
- RETCAMDocument42 paginiRETCAMRoberto Vega FloresÎncă nu există evaluări
- Cutting Back On Melanoma Biopsies: MelafindDocument5 paginiCutting Back On Melanoma Biopsies: MelafindAAKASHÎncă nu există evaluări
- Analysis of Skin Cancer Image Processing Using MATLABDocument5 paginiAnalysis of Skin Cancer Image Processing Using MATLABOmar MragÎncă nu există evaluări
- Malignancies of The Groin-Surgical and Anatomic Considerations-2018 PDFDocument248 paginiMalignancies of The Groin-Surgical and Anatomic Considerations-2018 PDFvirusÎncă nu există evaluări