Documente Academic
Documente Profesional
Documente Cultură
LNA-PNA Comparison4
Încărcat de
biosynthesis12Drepturi de autor
Formate disponibile
Partajați acest document
Partajați sau inserați document
Vi se pare util acest document?
Este necorespunzător acest conținut?
Raportați acest documentDrepturi de autor:
Formate disponibile
LNA-PNA Comparison4
Încărcat de
biosynthesis12Drepturi de autor:
Formate disponibile
LNA and PNA
probes
Comparison
612 East Main Street
Lewisville, TX 75057
EMAIL: biosyn@biosyn.com
FAX: 972-420-0442
PHONE: 972-420-8505
Toll Free: 1-800-227-0627
Introduction
Detection of nucleic acid sequences is widely used in many biological assays, with increasing penetration into clinical sciences
and other areas of life sciences. Quite often fluorescent tagged DNA sequences, either from naturally occurring sequences or
synthetic oligonucleotides, are used to probe a sample of interest. Synthetic non-natural analogs have also been found useful in
increasing specificity of detection. Here we take a look at two synthetic analogs: Locked Nucleic Acids (LNAs) and Peptide Nucleic
Acids (PNAs).
Thermodynamic comparison of LNA and PNA probes
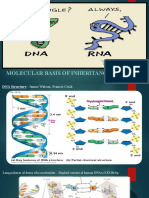
Structure of LNA
LNAs are a class of conformationally restricted nucleotide analogs, in which the
ribose ring is constrained by a methylene linkage between the 2’oxygen and
the 4’ carbon resulting in the locked 3’endo formation.(figure1) (Saenger,
1984).
The incorporation of LNA in an oligonucleotide increases the affinity of that
oligonucleotide for its complementary RNA or DNA target by increasing the
melting temperature (Tm) of the duplex. Additionally, the Tm difference between
a perfectly matched target and a mismatched target is substantially higher than
that observed when a DNA-based oligonucleotide is used. The properties like
—high Tm and excellent mismatch discrimination—make LNA-modified probes
ideal for analysis of short and very similar targets like miRNAs. Furthermore, by
adjusting the LNA content and probe length, it is possible to design Tm-normalized probes, thereby allowing hybridization condi-
tions that are optimal for all probes used on, for example, an array.
LNAs were seen to display increased thermodynamic stability and enhanced
nucleic acid recognition. Initial investigations of LNA melting temperatures (Tm) by Koshkin et al. (1998) revealed increased values
per LNA monomer(Tm) of +3 to +5oC and +4 to +8oC against complementary DNA and RNA oligonucleotides respectively.
Structural studies have shown that LNA oligonucleotides force an A-type (RNA-like) duplex conformation. Their melting tempera-
tures are increased by 1 to 8°C against DNA and 2 to 10°C against RNA by one LNA substitution in an oligonucleotide. An increase
in the number of LNA bases within an oligo, such as a qPCR probe increases Tm value significantly as can be seen below
Probe Sequence (5' ->3') LNA base Tm* Tm Tm/LNA
dG dT dG dA dT dA dT dG dC - 29ºC - -
dG + T dG dA + T dA +T dG dC 3 44ºC 15ºC +5ºC
+G+T+G+A+T+A+T+G+C 9 64ºC 35ºC +3.9ºC
Tm of duplex between probe and its complementary DNA sequence
Note: + symbol denotes the LNA base.
Structure of PNA
Peptide nucleic acids (PNAs) are oligonucleotide analogues in which the sugar-
phosphate backbone has been replaced by a pseudopeptide skeleton. They bind
DNA and RNA with high specificity and selectivity, leading to PNA–RNA and PNA-
DNA hybrids more stable than the corresponding nucleic acid complexes.
The thermal stability of PNA:DNA and PNA:RNA is higher compared to DNA:DNA
and DNA:RNA duplexes. The stronger binding is attributed to the lack of charge
repulsion between the neutral PNA and the negatively charged DNA or RNA strand
and also the PNAs are insensitive to variations in salt concentration and PNA:DNA
exhibit minor dependence from 10-50mM sodium chloride. This is in contrast to
DNA:DNA duplexes, which are highly dependent on ion concentration. At low
concentration PNA can bind to a target sequence at temperatures where DNA:DNA
hybridization is suboptimal.
Ability of PNA probes to invade double stranded DNA
PNA probes can bind to either single stranded DNA or RNA or to double stranded DNA
Homopyrimidine PNAs with a minimum of 10-mers, as well as PNAs containing a high proportion of pyrimidine
residues, bind to complementary DNA sequences to form highly stable (PNA)2–DNA triplex helices displaying Tm
over 70°C. In these triplexes, one PNA strand hybridizes to DNA through standard Watson–Crick base pairing
rules, while the other PNA strand binds to DNA through Hoogsteen hydrogen bonds. The resulting structure is
called P-loops. The stability of these triple helixes is so high that homopyrimidine PNA targeted to purine tracts of
dsDNA invades the duplex by displacing one of the DNA strands. The efficiency of this strand invasion can be
further enhanced by using two homopyrimidine PNA oligomers connected via a flexible linker or by the presence
of nonstandard nucleobases in the PNA molecule.
Stable triplex invasion complex, leading to the displacement of the second DNA strand into a 'D-loop'.
Significant applications of LNA probes
Application of LNA probes in miRNA analysis
miRNA analysis imposes many challenges, for example LNA modified capture probes may result in a more sensitive detection of miRNA in compari-
son to unmodified DNA-based capture probes. Furthermore, some miRNAs differ from each other by as little as a single nucleotide, emphasizing
the importance of good mismatch discrimination. LNAs have proven to be extremely useful in overcoming these obstacles, particularly in the areas
of miRNA profiling and in situ hybridization.
LNA fluorescent probes for Real Time (quantitative) PCR
Development of LNA fluorescent probes are an alternative in real time PCR and analytical assays including gene expression profiling, mutation
detection, allelic discrimination and pathogen detection
siLNA
LNA oligonucleotides and siLNA duplexes can be transfected into cells using standard reagents and have been shown to predictably mediate gene
silencing as single stranded antisense agents. Animal studies have confirmed the potency of single stranded antisense LNA oligonucleotides and
siLNA duplexes for gene silencing applications. Single stranded LNA antisense molecules are in clinical trials as anticancer agents
Applications of peptide nucleic acid probes
1) PNA fluorescent insitu hybridisation-
The PNA-FISH technique was first developed for quantitative telomere analysis. Using a unique fluorescein-labelled PNA
probe thus one can map the sequence onto a chromosome.
In FISH applications PNA probes have several remarkable properties. One of the most powerful aspects of PNA-probe based
explorations of chromosome structure is due to the exquisite base-discrimination in FISH applications. Very short PNA oligom-
ers can be effectively utilized. Despite the short length of the PNA oligomers the strength of interaction with target, and hence
the strength of the FISH signal, is greater than for a DNA oligomer. As a result PNA oligomer probes are capable of very
specific and discriminate interactions with nucleic acid targets. Research has shown that PNA probes are capable of single
base discrimination in FISH experiments. Because of the exquisite sequence discrimination of PNA probes in FISH applications PNA-FISH has
great potential in the growing area of Single Molecule Genomics
PNA may be used in many of the applications where PNA become a versatile tool in genetic diagnostics and a variety of molecular biology
techniques such as PCR clamping, plasmid vector tagging and nucleic acid solution phase hybridization detection
2) Antisense and antigene applications
The unique properties of PNA as a DNA mimic have been attributed for gene therapy and drug design sincePNAs can inhibit transcription and
translation of genes by tight binding to DNA or mRNA. For Eg: PNA mediated Transcription inhibition was demonstrated for the alpha-chain of the
interleukin-2 receptor, even an 8-mer PNA can efficiently block transcriptional elongation by (PNA)2–DNA triplex formation (Nielsen et al)
The mechanism of antisense activity has been attributed by RNase-H mediated degradation of the mRNA-oligonucleotide hybrid. Since PNAs are
not substrates for RNase, their antisense effect acts through steric interference of either RNA processing, transport into cytoplasm or translation,
caused by binding to the mRNA.
3) PNA as a major Delivery agent
PNAs can be use as adapters to link peptides, drugs or molecular tracers to plasmid vectors. According to the binding site, the
coupling of PNAs to plasmids has no effect either on the transcription of genes included in the plasmid or on the plasmid's physi-
ological activities. Thus, this approach allows circumvention of such barriers to gene transfer and fixation of drugs to plasmids in
order to enhance the gene delivery or tissue-specific targeting.
Using a triplex forming PNA as linker there is an eight times higher nuclear localization of a coupled nuclear localization signal
(NLS) than with the free oligonucleotide, (Braden et al).
General applications of LNA and PNA probes
1 LNA modified oligonucleotides are used as an excellent probes in FISH, The efficiency of PNAs as hybridization probes has also been demonstrated
combining high binding affinity with short hybridization time in fluorescence in situ hybridization (FISH) applications
2 LNA probes demonstrated superior allelic discrimination hence they PNA probes are used for the detection of single-nucleotide polymorphisms
have application in genotyping assays
3 LNA probes are used for hybridization studies or as real time polymerase LightUp and LightSpeed probes are unique reagents for real-time PCR,
chain reaction probes in the form of Taqman probes which combine fluorescent enhancement with the high sensitivity and
4 Chimeric LNA-DNA antisense constructs make them desirable for a host PNAs constitute as efficient compounds for effective antisense/antigene
of molecular applications application where PNA inhibits expression differently than antisense
oligonucleotides acting through RNase-H mediated degradation of the
mRNA–oligonucleotide hybrid
Pros and cons of LNA over PNA
LNA has higher affinity of hybridization than PNA(by up to 10°C substitu- Unmodified PNA cannot serve as a primer for polymerization, nor can it be
tion). a substrate for exonuclease activities of Taq polymerase.
LNA has a high aqueous solubility Whereas PNA has a low aqueous solubility
LNA-DNA chimeras can be easily synthesized in the lab as it is based on PNA on the other hand resembles closely on the peptide chemistry.
phosphoramidite chemistry Therefore PNA-DNA chimeras are not readily synthesized
Conclusion
PNA and LNA probes contributes in many areas of applications like antigene and antisense therapeutics, SNP genotyping assays,
Fluorescent insitu hybridization etc. Apart from the applications there are certain drawbacks where LNA probes are preferred over
PNA probes- firstly, PNA resembles closely on the peptide chemistry and therefore PNA-DNA chimeras are not readily synthe-
sized where as LNAs are synthesized using conventional phosphoramidite chemistry.
LNA substitution couple the high affinity binding that characterize recognition by PNAs with retention of phosphodiester backbone
of DNA or RNA. Therefore it is supposed that LNA’s would be ideal candidates for examining whether efficient inhibition of gene
expression could be extended beyond PNAs to negatively charged, synthetic oligonucleotides.
References
1) Bondensgaard, K., M. Petersen, S. K. Singh, V. K. Rajwanshi, R. Kumar, J. Wengel, and J. P. Jacobsen. 2000. Structural stud-
ies of LNA:RNA duplexes by NMR: conformations and implications for RNase H activity. Chemistry 6:2687-2695.
2)Egholm, M., O. Buchardt, L. Christensen, C. Behrens, S. M. Freier, D. A. Driver, R. H. Berg, S. K. Kim, B. Norden, and P. E.
Nielsen. 1993.
3) Perry-O'Keefe, H., S. Rigby, K. Oliveira, D. Sørensen, H. Stender, J. Coull, and J. J. Hyldig-Nielsen. 2001. Identification of
indicator microorganisms using a standardized PNA FISH method. J. Microbiol. Methods 47:281-292
4) McTigue, P.M.; Peterson, R.J.; Kahn, J.D. Sequence-dependent thermodynamic parameters; for locked nucleic acid (LNA)-
DNA duplex formation. Biochemistry. 2004;43:5388–5405. [PubMed]
5) Castoldi, M. et al. A sensitive array for micro RNA expression profiling (miChip) based on locked nucleic acids (LNA). RNA
6)Fiandaca, M.J.; Hyldig-Nielsen, J.J.; Gildea, B.D.; Coull,J.M. Self-reporting PNA/DNA primers for PCR analysis.Genome Res.
2001, 11, 609–613
S-ar putea să vă placă și
- RNA Methodologies: A Laboratory Guide for Isolation and CharacterizationDe la EverandRNA Methodologies: A Laboratory Guide for Isolation and CharacterizationÎncă nu există evaluări
- PCR in Infectious DiseasesDocument3 paginiPCR in Infectious Diseasesthị sô phiaÎncă nu există evaluări
- The Peptide Nucleic Acids (Pnas) : A New Generation of Probes For Genetic and Cytogenetic AnalysesDocument10 paginiThe Peptide Nucleic Acids (Pnas) : A New Generation of Probes For Genetic and Cytogenetic AnalysesEhsan HumayunÎncă nu există evaluări
- DNA in Supramolecular Chemistry and NanotechnologyDe la EverandDNA in Supramolecular Chemistry and NanotechnologyEugen StulzÎncă nu există evaluări
- Subject Name: Genetic Engineering-II Subject Code: MTI-605 Unit No:5 Unit Name: Molecular DiagnosticsDocument13 paginiSubject Name: Genetic Engineering-II Subject Code: MTI-605 Unit No:5 Unit Name: Molecular DiagnosticsSuman DeshmukhÎncă nu există evaluări
- Peptide Nucleic AcidDocument27 paginiPeptide Nucleic AcidDharmendra KumarÎncă nu există evaluări
- DNA Biosensors Based On Peptide Nucleic Acid PNA Recognition Layers A Review1 - 1998 - Biosensors and BioelectronicsDocument6 paginiDNA Biosensors Based On Peptide Nucleic Acid PNA Recognition Layers A Review1 - 1998 - Biosensors and BioelectronicswardaninurindahÎncă nu există evaluări
- Mrs. Tole S.B. Asst. Prof. K.T.Patil College of Pharmacy, OsmanabadDocument22 paginiMrs. Tole S.B. Asst. Prof. K.T.Patil College of Pharmacy, OsmanabadShobha Tole100% (2)
- Habb - HBB 231 Notes - June 2024Document196 paginiHabb - HBB 231 Notes - June 2024AmandaÎncă nu există evaluări
- Submitted By: Akshat Goel Xii - C R.NoDocument17 paginiSubmitted By: Akshat Goel Xii - C R.NoAkshat GoelÎncă nu există evaluări
- PCR, RT-PCR and QPCR: Dr. Sandeep Agrawal MD Senior Resident & PHD Scholar Department of Biochemistry Aiims, New DelhiDocument35 paginiPCR, RT-PCR and QPCR: Dr. Sandeep Agrawal MD Senior Resident & PHD Scholar Department of Biochemistry Aiims, New DelhiDanyÎncă nu există evaluări
- Polymerase Chain Reaction (PCR)Document67 paginiPolymerase Chain Reaction (PCR)Derick SemÎncă nu există evaluări
- Genetic Control: Dna and RnaDocument18 paginiGenetic Control: Dna and Rnarasha nadaÎncă nu există evaluări
- NCERT Solutions For Class 12 Biology Chapter 6 Molecular Basis of InheritanceDocument10 paginiNCERT Solutions For Class 12 Biology Chapter 6 Molecular Basis of InheritanceSneha HipparkarÎncă nu există evaluări
- Rev. Lect 2. MOLECULAR TECHNIQUES IN DIAGNOSTIC MICROBIOLOGYDocument75 paginiRev. Lect 2. MOLECULAR TECHNIQUES IN DIAGNOSTIC MICROBIOLOGYKhrys HardyÎncă nu există evaluări
- Nucleic Acids SummaryDocument4 paginiNucleic Acids SummaryIvan MiglioriniÎncă nu există evaluări
- PCR1Document4 paginiPCR1Hina NoorÎncă nu există evaluări
- Polymerase Chain Reaction ProtocolDocument14 paginiPolymerase Chain Reaction ProtocolDespoina ChatziÎncă nu există evaluări
- PCRDocument3 paginiPCRarian.boostani2002Încă nu există evaluări
- Polymerase Chain Reaction (PCR)Document17 paginiPolymerase Chain Reaction (PCR)Nasir MahmudÎncă nu există evaluări
- pcr شرح مفصل عنDocument26 paginipcr شرح مفصل عنZainab HasanÎncă nu există evaluări
- Param's PCR ProjectDocument11 paginiParam's PCR ProjectPARRU xYTÎncă nu există evaluări
- Activity 4.1.2 PCR WorksheetDocument3 paginiActivity 4.1.2 PCR Worksheet3021031Încă nu există evaluări
- Dna Analysis - 1.2Document21 paginiDna Analysis - 1.2Aparna PrahladÎncă nu există evaluări
- PCR: An Outstanding MethodDocument17 paginiPCR: An Outstanding MethodPriyaranjan KumarÎncă nu există evaluări
- Ritobrata Goswami: Email: Tel: 03222-284570Document44 paginiRitobrata Goswami: Email: Tel: 03222-284570Ashmit RanjanÎncă nu există evaluări
- Prinsip PCRDocument4 paginiPrinsip PCRZaid al GifariÎncă nu există evaluări
- Polymerase Chain Reaction Pre-LabDocument9 paginiPolymerase Chain Reaction Pre-LabChristian GalangÎncă nu există evaluări
- Target Amplification MethodsDocument3 paginiTarget Amplification MethodsNOR-FATIMAH BARATÎncă nu există evaluări
- DR Tiang Report 1Document7 paginiDR Tiang Report 1KhAi EnÎncă nu există evaluări
- DNA Renaturation & HybridizationDocument3 paginiDNA Renaturation & HybridizationAqsa RaffaqÎncă nu există evaluări
- Lect 7Document18 paginiLect 7ayeshaÎncă nu există evaluări
- Methods Molecular Cardiology:n: The Polymerase Chain ReactionDocument7 paginiMethods Molecular Cardiology:n: The Polymerase Chain ReactionmarcelsÎncă nu există evaluări
- DNA STRUCTURE LectureDocument44 paginiDNA STRUCTURE Lectureevacarlina1721Încă nu există evaluări
- تقرير مايكروDocument1 paginăتقرير مايكروEzz WlsmanÎncă nu există evaluări
- Praxi LabDocument17 paginiPraxi LabRetro GirlÎncă nu există evaluări
- Bio Project-Polymerase Chain ReactionDocument21 paginiBio Project-Polymerase Chain ReactionS.AbiniveshÎncă nu există evaluări
- Genomics 2Document15 paginiGenomics 21SI19BT002 AVINASH K AÎncă nu există evaluări
- Northern Blot PDFDocument8 paginiNorthern Blot PDFerick ortiz lopezÎncă nu există evaluări
- FragmentDocument1 paginăFragmentGheorghe Georq NitescuÎncă nu există evaluări
- Molecular BiologyDocument2 paginiMolecular BiologyJaymark HisonaÎncă nu există evaluări
- Mol Basis of InheritanceDocument74 paginiMol Basis of InheritanceNishita BharaliÎncă nu există evaluări
- Chapter 5 MurrayDocument3 paginiChapter 5 MurrayTotalenesya Reforrent SutiknoÎncă nu există evaluări
- Polymerase Chain ReactionDocument2 paginiPolymerase Chain ReactionՏᗅႮRᗅBH ՏᗅiNiÎncă nu există evaluări
- ICS 2 Notes: 1. Gel ElectrophoresisDocument6 paginiICS 2 Notes: 1. Gel ElectrophoresisTan Yu YangÎncă nu există evaluări
- Genetics Engineering PPT Grp#09Document52 paginiGenetics Engineering PPT Grp#09Alina RajputÎncă nu există evaluări
- Science of Living System: Ritobrata GoswamiDocument49 paginiScience of Living System: Ritobrata GoswamiDeath TromeÎncă nu există evaluări
- MTPC 140: Molecular Biology and DiagnosticsDocument26 paginiMTPC 140: Molecular Biology and DiagnosticsValdez Francis ZaccheauÎncă nu există evaluări
- Contamination Proof-Reading: Mplification Ensitivity Pecificity Obustness Utomation Ell-Free Method Ase of Set-UpDocument4 paginiContamination Proof-Reading: Mplification Ensitivity Pecificity Obustness Utomation Ell-Free Method Ase of Set-UpSheila ChaiÎncă nu există evaluări
- 6 Molecular Genetics Part IIDocument51 pagini6 Molecular Genetics Part II2021-204943Încă nu există evaluări
- Review Approaches To DNA Mutagenesis: An Overview: Michael Mingfu Ling and Brian H. RobinsonDocument22 paginiReview Approaches To DNA Mutagenesis: An Overview: Michael Mingfu Ling and Brian H. RobinsonIngeniería QuímicaÎncă nu există evaluări
- Exercise 8 Polymerase Chain Reaction: Cell and Molecular Biology LaboratoryDocument7 paginiExercise 8 Polymerase Chain Reaction: Cell and Molecular Biology LaboratoryDham DoñosÎncă nu există evaluări
- LECT-12 The Polymerase Chain Reaction (20192020-1)Document23 paginiLECT-12 The Polymerase Chain Reaction (20192020-1)Ikrimah FithriyandiniÎncă nu există evaluări
- TRANSCRIPTIONDocument1 paginăTRANSCRIPTIONLouise Mae LolorÎncă nu există evaluări
- Laboratory TechniquesDocument50 paginiLaboratory TechniquesmÎncă nu există evaluări
- DNA Kita Iiwan RNA CytoDocument3 paginiDNA Kita Iiwan RNA CytozairahÎncă nu există evaluări
- Final DGDocument9 paginiFinal DGSulav AcharyaÎncă nu există evaluări
- What Is PCR?: T. Aquaticus Lives in Hot Springs and Hydrothermal Vents. Its DNA Polymerase Is Very Heat-StableDocument3 paginiWhat Is PCR?: T. Aquaticus Lives in Hot Springs and Hydrothermal Vents. Its DNA Polymerase Is Very Heat-StableFaiz RasoolÎncă nu există evaluări
- Biotech Group 5Document6 paginiBiotech Group 5CarminexxxÎncă nu există evaluări
- HLA Analysis Poster With LegendDocument1 paginăHLA Analysis Poster With Legendbiosynthesis12Încă nu există evaluări
- Phagocytosis in A Chemr23-And Chemerin Peptides Promote: Data SupplementaryDocument11 paginiPhagocytosis in A Chemr23-And Chemerin Peptides Promote: Data Supplementarybiosynthesis12Încă nu există evaluări
- Peptides As Molecular Imaging ProbesDocument3 paginiPeptides As Molecular Imaging Probesbiosynthesis12Încă nu există evaluări
- Direct, Electronic Microrna Detection For The Rapid Determination of Differential Expression ProfilesDocument4 paginiDirect, Electronic Microrna Detection For The Rapid Determination of Differential Expression Profilesbiosynthesis12Încă nu există evaluări
- User Bulletin #2: Abi P 7700 Sequence Detection SystemDocument36 paginiUser Bulletin #2: Abi P 7700 Sequence Detection Systembiosynthesis12Încă nu există evaluări
- TaqMan Universal PCR Master MixDocument60 paginiTaqMan Universal PCR Master Mixbiosynthesis12Încă nu există evaluări
- DNA/RNA Real-Time Quantitative PCRDocument8 paginiDNA/RNA Real-Time Quantitative PCRbiosynthesis12Încă nu există evaluări
- Alexa FluorDocument2 paginiAlexa Fluorbiosynthesis12Încă nu există evaluări
- Probes and Primer Design Using Primer ExpressDocument15 paginiProbes and Primer Design Using Primer Expressbiosynthesis12Încă nu există evaluări
- Peptide MsdsDocument2 paginiPeptide Msdsbiosynthesis12Încă nu există evaluări
- Peptide MsdsDocument2 paginiPeptide Msdsbiosynthesis12Încă nu există evaluări
- Workshop PresentationDocument24 paginiWorkshop Presentationhuang SteffiÎncă nu există evaluări
- MolecularDocument8 paginiMolecularHassanAl-shaweshÎncă nu există evaluări
- Genes AssignmentDocument4 paginiGenes AssignmentKyle Hilary MatundingÎncă nu există evaluări
- The Central Dogma of Molecular BiologyDocument45 paginiThe Central Dogma of Molecular BiologyEirla Kneriz Trinidad67% (3)
- Submitted by Aiswarya V 1St MSC Zoology Roll Number 3301Document32 paginiSubmitted by Aiswarya V 1St MSC Zoology Roll Number 3301PES PEOPLEÎncă nu există evaluări
- Protein Synthesis: Indian Institute of Technology PatnaDocument29 paginiProtein Synthesis: Indian Institute of Technology PatnaHritik KumarÎncă nu există evaluări
- Kami Export - Worksheet 14 Central DogmaDocument2 paginiKami Export - Worksheet 14 Central DogmaLindsey TamlinÎncă nu există evaluări
- Nitrogenous BasesDocument6 paginiNitrogenous BasesMaak Ali AnsariÎncă nu există evaluări
- IB Biology Notes - 35 Transcription TranslationDocument2 paginiIB Biology Notes - 35 Transcription TranslationJohn Philip D. NapalÎncă nu există evaluări
- Dna Replication: Meselson & Stahl's Experiment (1958)Document4 paginiDna Replication: Meselson & Stahl's Experiment (1958)Angie LawrenceÎncă nu există evaluări
- BCH 323 NotesDocument41 paginiBCH 323 NotesvictorÎncă nu există evaluări
- Division of Capiz: Capiz@deped - Gov.phDocument7 paginiDivision of Capiz: Capiz@deped - Gov.phRONALD ARTILLEROÎncă nu există evaluări
- cDNA LibrariesDocument26 paginicDNA LibrariesRavi DesaiÎncă nu există evaluări
- Test Nucleic Acids SLDocument2 paginiTest Nucleic Acids SLKatrīna SimanovskaÎncă nu există evaluări
- RiboswitchesDocument30 paginiRiboswitchesYaseswini NeelamrajuÎncă nu există evaluări
- DNA HistoryDocument21 paginiDNA HistoryGlenn Valentin MendozaÎncă nu există evaluări
- FUNDBIO Central Dogma ActivityDocument3 paginiFUNDBIO Central Dogma ActivityViktor SolomonÎncă nu există evaluări
- Chapter 2 HomeworkDocument5 paginiChapter 2 HomeworkKvn4N6Încă nu există evaluări
- Difference Between Dna & Rna: Students NameDocument5 paginiDifference Between Dna & Rna: Students NameHisham MustafaÎncă nu există evaluări
- Science: Quarter 3 - Module 4.1: Protein SynthesisDocument32 paginiScience: Quarter 3 - Module 4.1: Protein SynthesisMariiÎncă nu există evaluări
- Unit 12 - DNA Worksheet - Structure of DNA and ReplicationDocument4 paginiUnit 12 - DNA Worksheet - Structure of DNA and ReplicationRibbitÎncă nu există evaluări
- QuizDocument1 paginăQuizJanÎncă nu există evaluări
- Biological Significance of DNA and RNA SturucturesDocument2 paginiBiological Significance of DNA and RNA SturucturesDr.Charles BhaskarÎncă nu există evaluări
- Annotated-Kami Export - DNA WorksheetDocument6 paginiAnnotated-Kami Export - DNA WorksheetFredrick DanielsÎncă nu există evaluări
- TranscriptionDocument30 paginiTranscriptionAizelle TarataraÎncă nu există evaluări
- Binas 6e Druk 68BDocument1 paginăBinas 6e Druk 68BGwen Van de WallÎncă nu există evaluări
- Nucleotides and Nucleic AcidsDocument65 paginiNucleotides and Nucleic AcidsnutriÎncă nu există evaluări
- Transcription in Bacteria: E. ColiDocument3 paginiTranscription in Bacteria: E. ColiGuilliane GallanoÎncă nu există evaluări
- The Dna StructureDocument28 paginiThe Dna Structuremark gonzalesÎncă nu există evaluări
- Dna & RnaDocument43 paginiDna & RnaheyyyaleÎncă nu există evaluări
- 10% Human: How Your Body's Microbes Hold the Key to Health and HappinessDe la Everand10% Human: How Your Body's Microbes Hold the Key to Health and HappinessEvaluare: 4 din 5 stele4/5 (33)
- Masterminds: Genius, DNA, and the Quest to Rewrite LifeDe la EverandMasterminds: Genius, DNA, and the Quest to Rewrite LifeÎncă nu există evaluări
- Why We Die: The New Science of Aging and the Quest for ImmortalityDe la EverandWhy We Die: The New Science of Aging and the Quest for ImmortalityEvaluare: 4.5 din 5 stele4.5/5 (6)
- When the Body Says No by Gabor Maté: Key Takeaways, Summary & AnalysisDe la EverandWhen the Body Says No by Gabor Maté: Key Takeaways, Summary & AnalysisEvaluare: 3.5 din 5 stele3.5/5 (2)
- Tales from Both Sides of the Brain: A Life in NeuroscienceDe la EverandTales from Both Sides of the Brain: A Life in NeuroscienceEvaluare: 3 din 5 stele3/5 (18)
- The Rise and Fall of the Dinosaurs: A New History of a Lost WorldDe la EverandThe Rise and Fall of the Dinosaurs: A New History of a Lost WorldEvaluare: 4 din 5 stele4/5 (597)
- Buddha's Brain: The Practical Neuroscience of Happiness, Love & WisdomDe la EverandBuddha's Brain: The Practical Neuroscience of Happiness, Love & WisdomEvaluare: 4 din 5 stele4/5 (216)
- Return of the God Hypothesis: Three Scientific Discoveries That Reveal the Mind Behind the UniverseDe la EverandReturn of the God Hypothesis: Three Scientific Discoveries That Reveal the Mind Behind the UniverseEvaluare: 4.5 din 5 stele4.5/5 (52)
- A Brief History of Intelligence: Evolution, AI, and the Five Breakthroughs That Made Our BrainsDe la EverandA Brief History of Intelligence: Evolution, AI, and the Five Breakthroughs That Made Our BrainsEvaluare: 4.5 din 5 stele4.5/5 (6)
- The Molecule of More: How a Single Chemical in Your Brain Drives Love, Sex, and Creativity--and Will Determine the Fate of the Human RaceDe la EverandThe Molecule of More: How a Single Chemical in Your Brain Drives Love, Sex, and Creativity--and Will Determine the Fate of the Human RaceEvaluare: 4.5 din 5 stele4.5/5 (517)
- Gut: the new and revised Sunday Times bestsellerDe la EverandGut: the new and revised Sunday Times bestsellerEvaluare: 4 din 5 stele4/5 (393)
- Seven and a Half Lessons About the BrainDe la EverandSeven and a Half Lessons About the BrainEvaluare: 4 din 5 stele4/5 (111)
- Undeniable: How Biology Confirms Our Intuition That Life Is DesignedDe la EverandUndeniable: How Biology Confirms Our Intuition That Life Is DesignedEvaluare: 4 din 5 stele4/5 (11)
- All That Remains: A Renowned Forensic Scientist on Death, Mortality, and Solving CrimesDe la EverandAll That Remains: A Renowned Forensic Scientist on Death, Mortality, and Solving CrimesEvaluare: 4.5 din 5 stele4.5/5 (397)
- The Ancestor's Tale: A Pilgrimage to the Dawn of EvolutionDe la EverandThe Ancestor's Tale: A Pilgrimage to the Dawn of EvolutionEvaluare: 4 din 5 stele4/5 (812)
- Who's in Charge?: Free Will and the Science of the BrainDe la EverandWho's in Charge?: Free Will and the Science of the BrainEvaluare: 4 din 5 stele4/5 (65)
- Good Without God: What a Billion Nonreligious People Do BelieveDe la EverandGood Without God: What a Billion Nonreligious People Do BelieveEvaluare: 4 din 5 stele4/5 (66)
- Moral Tribes: Emotion, Reason, and the Gap Between Us and ThemDe la EverandMoral Tribes: Emotion, Reason, and the Gap Between Us and ThemEvaluare: 4.5 din 5 stele4.5/5 (116)
- Remnants of Ancient Life: The New Science of Old FossilsDe la EverandRemnants of Ancient Life: The New Science of Old FossilsEvaluare: 3 din 5 stele3/5 (3)
- The Lives of Bees: The Untold Story of the Honey Bee in the WildDe la EverandThe Lives of Bees: The Untold Story of the Honey Bee in the WildEvaluare: 4.5 din 5 stele4.5/5 (44)
- Darwin's Doubt: The Explosive Origin of Animal Life and the Case for Intelligent DesignDe la EverandDarwin's Doubt: The Explosive Origin of Animal Life and the Case for Intelligent DesignEvaluare: 4 din 5 stele4/5 (19)
- The Consciousness Instinct: Unraveling the Mystery of How the Brain Makes the MindDe la EverandThe Consciousness Instinct: Unraveling the Mystery of How the Brain Makes the MindEvaluare: 4.5 din 5 stele4.5/5 (93)
- Human: The Science Behind What Makes Your Brain UniqueDe la EverandHuman: The Science Behind What Makes Your Brain UniqueEvaluare: 3.5 din 5 stele3.5/5 (38)
- The Other Side of Normal: How Biology Is Providing the Clues to Unlock the Secrets of Normal and Abnormal BehaviorDe la EverandThe Other Side of Normal: How Biology Is Providing the Clues to Unlock the Secrets of Normal and Abnormal BehaviorÎncă nu există evaluări